Abstract
Mycoplasma ovipneumoniae (Mo) is difficult to culture, resulting in many difficulties in related research and application. Since nucleotide metabolism is a basic metabolism affects growth, this study conducted a “point-to-point” comparison of the corresponding growth phases between the Mo NM151 strain and the Mycoplasma mycoides subsp. capri (Mmc) PG3 strain. The results showed that the largest difference in nucleotide metabolism was found in the stationary phase. Nucleotide synthesis in PG3 was mostly de novo, while nucleotide synthesis in NM151 was primarily based on salvage synthesis. Compared with PG3, the missing reactions of NM151 referred to the synthesis of deoxythymine monophosphate. We proposed and validated a culture medium with added serine to fill this gap and prolong the stationary phase of NM151. This solved the problem of the fast death of Mo, which is significant for related research and application.
Go to : 
References
1. Wang XH, Wang YF, Huang HB, Bai F, Shi X, Ma C, Gao Y, Zhang J, Zhang W, Hao Y. Understanding the metabolism of Mycoplasma mycoides subsp. capri in vitro by a transcriptomic analysis. J Integr Agric. 2018; 17:428–435.

2. Razin S, Yogev D, Naot Y. Molecular biology and pathogenicity of mycoplasmas. Microbiol Mol Biol Rev. 1998; 62:1094–1156.

4. Wodke JA, Puchałka J, Lluch-Senar M, Marcos J, Yus E, Godinho M, Gutiérrez-Gallego R, dos Santos VA, Serrano L, Klipp E, Maier T. Dissecting the energy metabolism in Mycoplasma pneumoniae through genome-scale metabolic modeling. Mol Syst Biol. 2013; 9:653.

5. Yus E, Maier T, Michalodimitrakis K, van Noort V, Yamada T, Chen WH, Wodke JA, Güell M, Martínez S, Bourgeois R, Kühner S, Raineri E, Letunic I, Kalinina OV, Rode M, Herrmann R, Gutiérrez-Gallego R, Russell RB, Gavin AC, Bork P, Serrano L. Impact of genome reduction on bacterial metabolism and its regulation. Science. 2009; 326:1263–1268.

6. Wang XH, Huang HB, Cheng C, Wang RC, Zheng JQ, Hao YQ, Zhang W. Complete genome sequence of Mycoplasma ovipneumoniae strain NM2010, which was isolated from a sheep in China. J Integr Agric. 2014; 13:2562–2563.

7. Xu CG, Hao YQ, Zhang L, Hao RX, Liu XL, Huang ZY. Molecular cloning and immune response analysis of putative variable lipoproteins from Mycoplasma mycoides subsp capri. Genet Mol Res. 2014; 13:1527–1539.

9. McClure R, Balasubramanian D, Sun Y, Bobrovskyy M, Sumby P, Genco CA, Vanderpool CK, Tjaden B. Computational analysis of bacterial RNA-Seq data. Nucleic Acids Res. 2013; 41:e140.

10. Sanavia T, Finotello F, Di Camillo B. FunPat: function-based pattern analysis on RNA-seq time series data. BMC Genomics. 2015; 16:S2.

11. Kanehisa M, Goto S, Hattori M, Aoki-Kinoshita KF, Itoh M, Kawashima S, Katayama T, Araki M, Hirakawa M. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 2006; 34:D354–D357.

12. Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2008; 36:D480–D484.

13. Sun S, Xuan F, Ge X, Fu H, Zhu J, Zhang S. Identification of differentially expressed genes in hepatopancreas of oriental river prawn, Macrobrachium nipponense exposed to environmental hypoxia. Gene. 2014; 534:298–306.

14. Ziemkowski P, Felczak K, Poznański J, Kulikowski T, Zieliński Z, Cieś la J, Rode W. Interactions of 2′-fluoro-substituted dUMP analogues with thymidylate synthase. Biochem Biophys Res Commun. 2007; 362:37–43.

15. Wang JY. Biochemistry. Beijing: Higher Education Press;2002. 400 p.
Go to : 
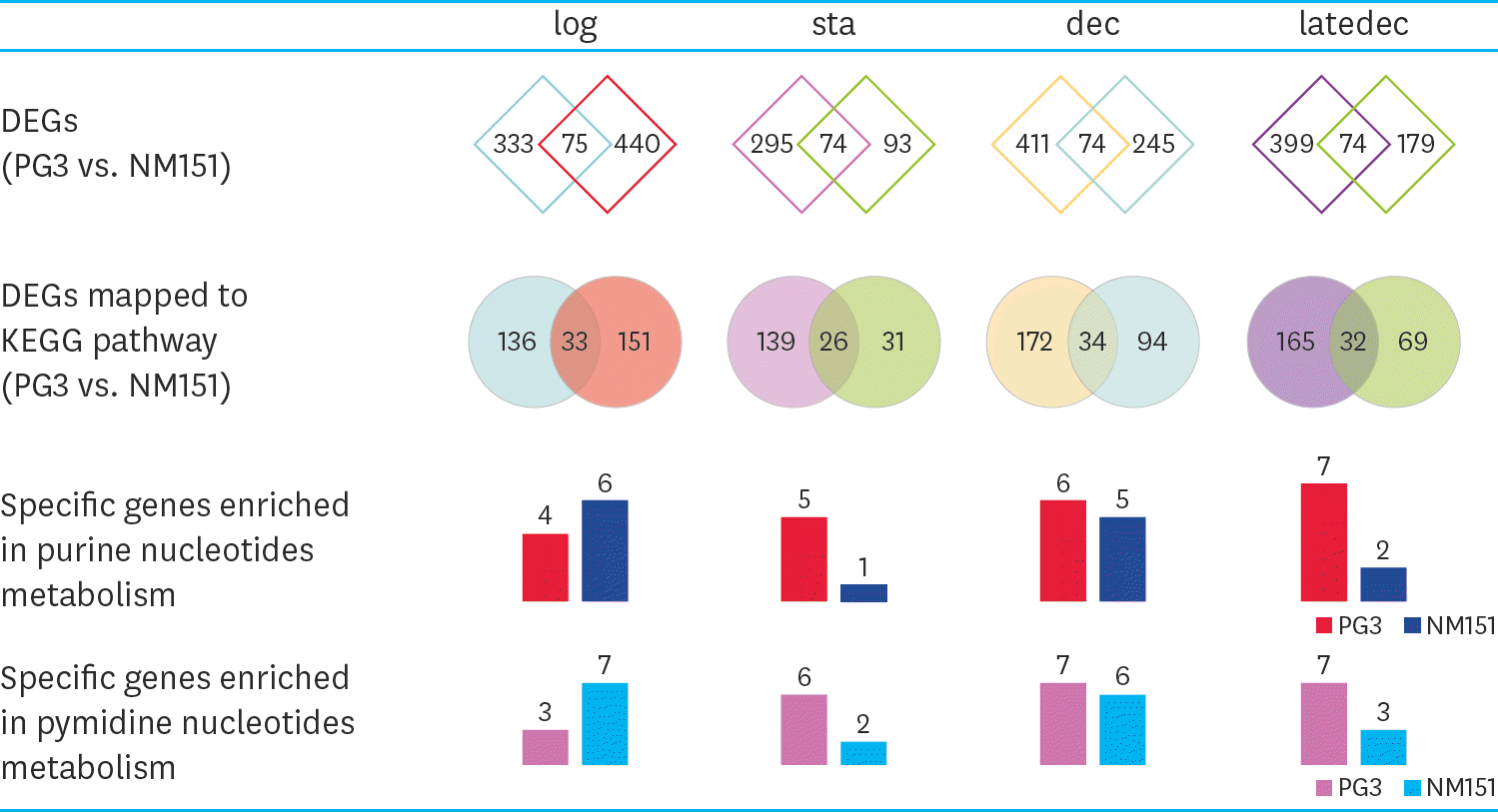 | Fig. 1.Statistics of DEGs per group (PG3 vs. NM151). The first line is the number of DEGs and core genes of PG3 and NM151 in the four comparison groups, respectively. Same as above, the second line is the number of DEGs and core genes enriched in the KEGG pathway. The third and fourth lines are the specific genes of PG3 and NM151 enriched in purine nucleotides and pymidine nucleotides metabolism. DEG, differentially expressed gene; KEGG, Kyoto Encyclopedia of Genes and Genomes |
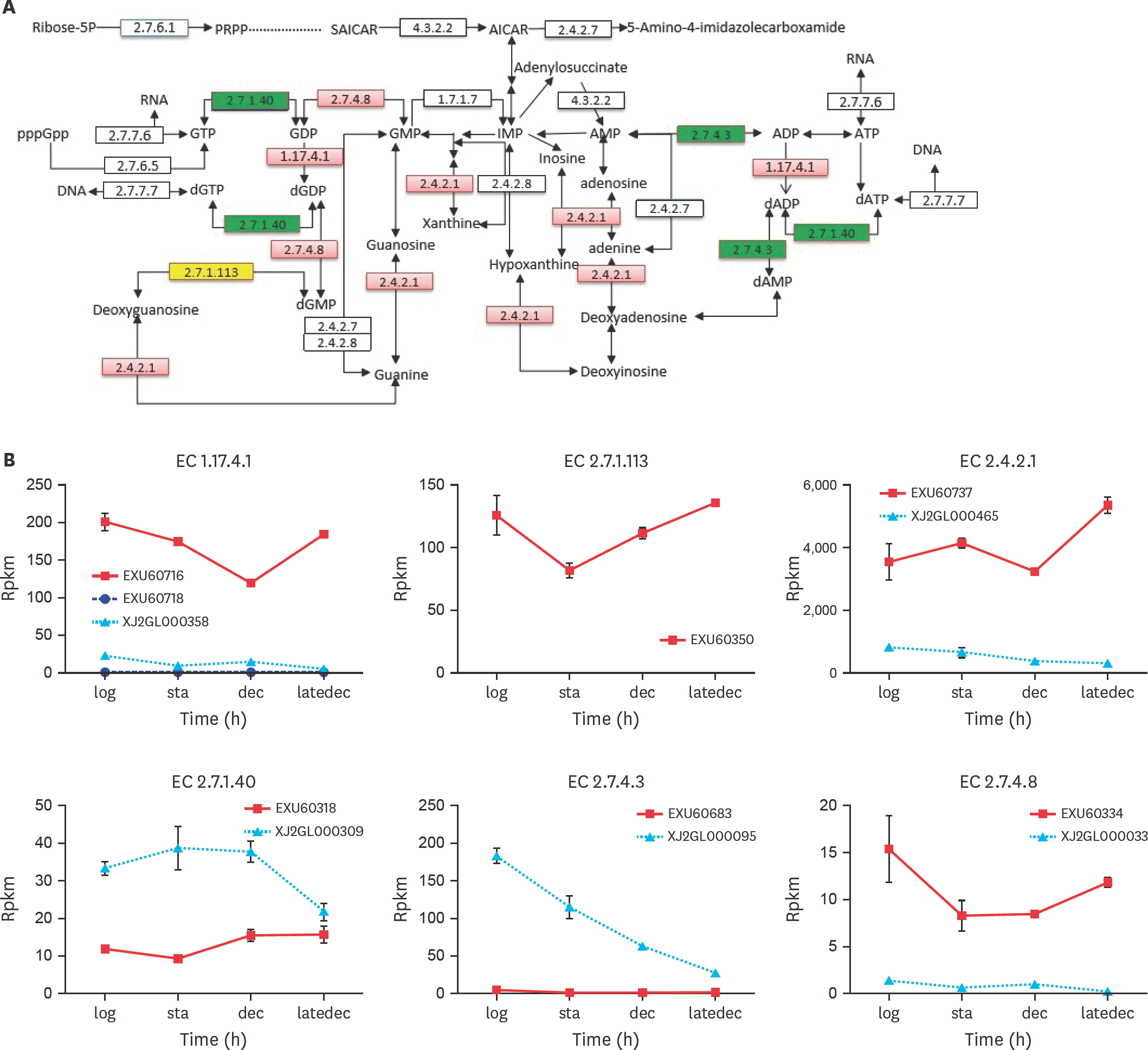 | Fig. 2.Specific enzymes in purine metabolism. (A) was a pathway part of Ko00230, only the enzymes, whose differentially expressed genes were coded between the two species, were listed. The enzyme boxes were divided into four equal parts, one for each growth phase. If the gene were upregulaed in PG3 compared with NM151, this enzyme rectangle in red. And the opposite (downregulation) could be green. If the gene is not expressed in NM151, it will be yellow. (B) were the RPKM value of genes in PG3 and NM151 during the four growth stages. RPKM, reads per kilobase per million mapped; PRPP, Phosphoribosyl pyrophosphate; SAICAR, phosphoribosylaminoimidazolesuccinocarboxamide; AICAR, aminoimidazole carboxamide ribonucleotide; GTP, guanosine triphosphate; GDP, guanosine diphosphate; GMP, guanosine monophosphate; IMP, inosine monophosphate; AMP, adenosine monophosphate; ADP, adenosine diphosphate; ATP, adenosine triphosphate; d, deoxy-. |
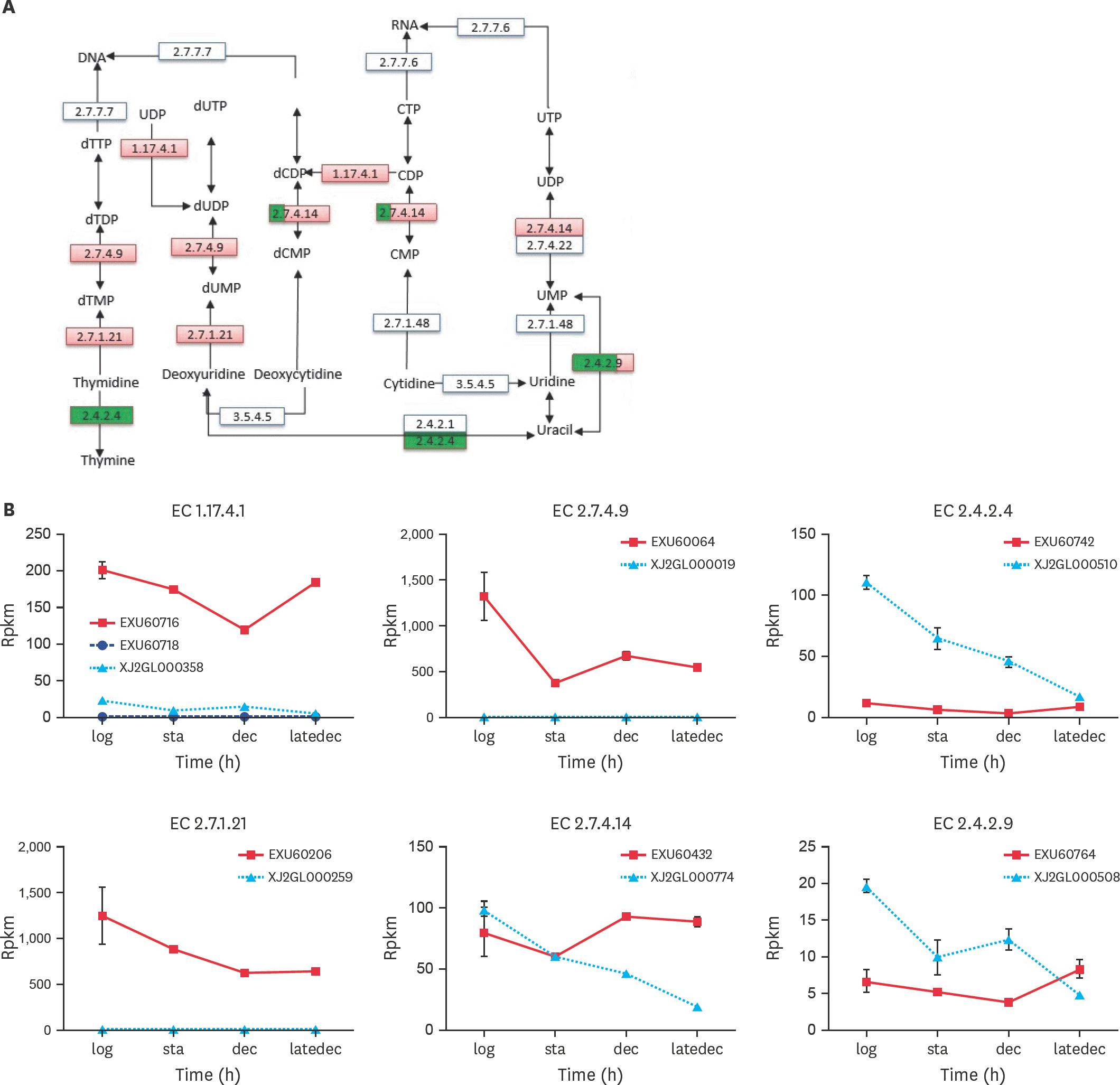 | Fig. 3.Specific enzymes in pyrimidine metabolism. (A) was a pathway part of Ko00240, only the enzymes, whose differentially expressed genes were coded between the 2 species, were listed. The enzyme boxes were divided into 4 equal parts, one for each growth phase. If the gene were upregulaed in PG3 compared with NM151, this enzyme rectangle in red. And the opposite (downregulation) could be green. If the gene is not expressed in NM151, it will be yellow. (B) were the RPKM value of genes in PG3 and NM151 during the 4 growth stages. TTP, thiamine triphosphate; TDP, thiamine diphosphate; TMP, thiamine monophosphate; UDP, uridine diphosphate; UTP, uridine triphosphate; UMP, uridine monophosphate; CTP, cytidine triphosphate; CMP, cytidine monophosphate; CDP, cytidine diphosphate; d, deoxy-. |
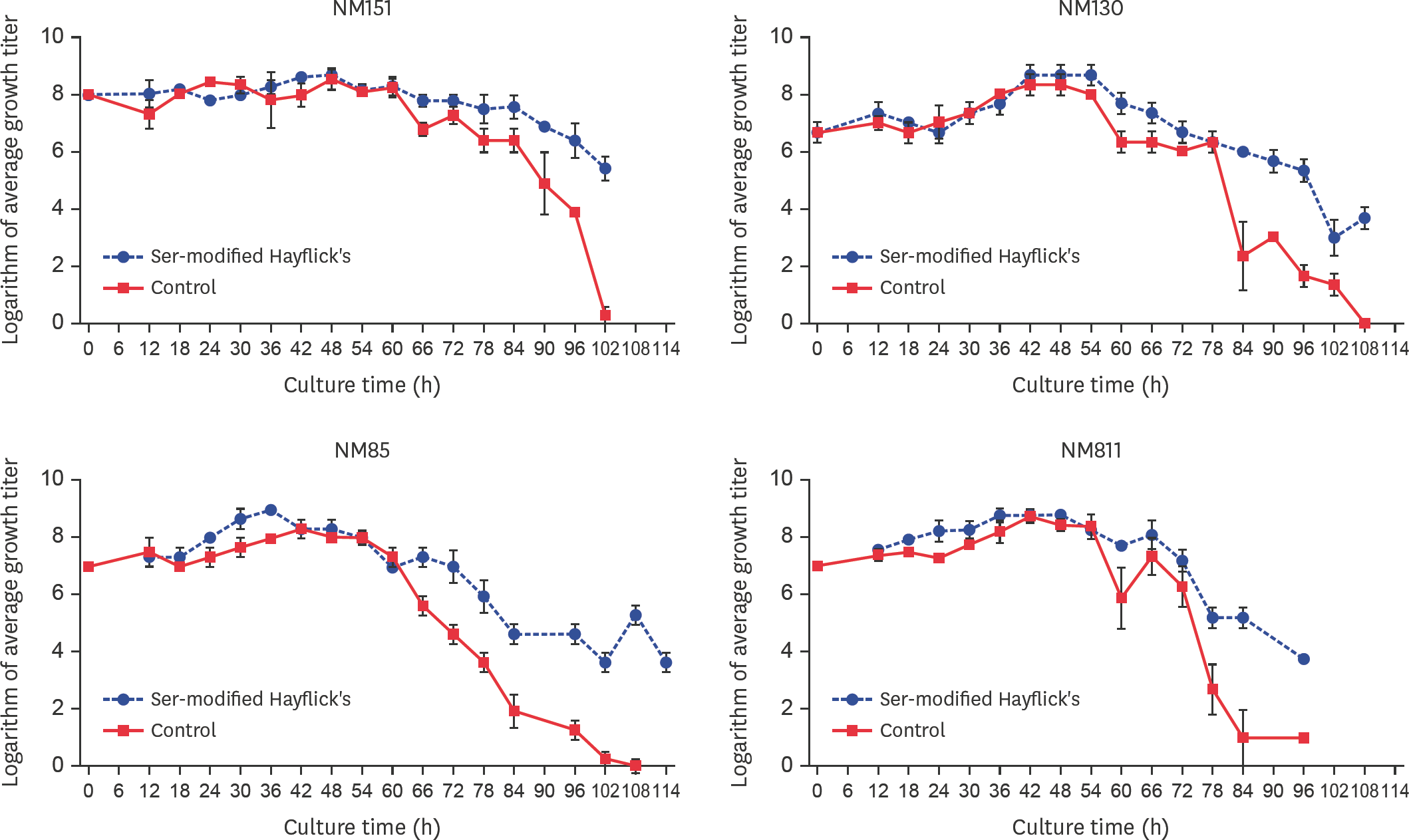 | Fig. 4.Growth curves in ser-modified Hayflick's mediums. All cultures were performed three times, and colonies were counted in duplicate for each dilution. The growth curve of the Mo NM151 strain was constructed by plotting the logarithm of the colony counts for each dilution on the Y-axis and the sampling time on the X-axis. |
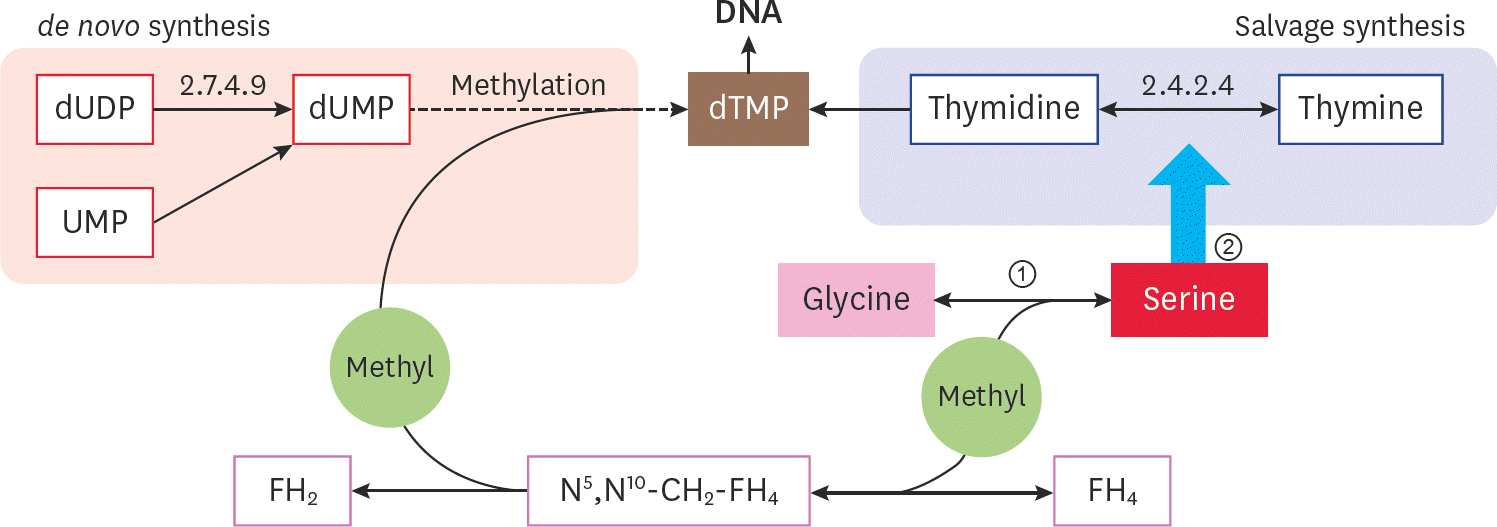 | Fig. 5.The pathway of serine participates in the metabolism of nucleotides. ① serine can provide methyl for dUMP converting to dTMP and contributes to the de novo synthesis of thymine nucleotides; ② mycoplasma can produce thymidine phosphorylase (EC 2.4.2.4) when consuming serine in the cell culture medium supernatant. |




 PDF
PDF Citation
Citation Print
Print


 XML Download
XML Download