Abstract
Background
Methods
Results
Notes
Author Contributions:
The contributions of the authors are as follows:
Research conception and design: Nakano R, Yano H.
Data acquisition: Kakuta N, Nakano R, Nakano A, Suzuki Y, Tanouchi A, Masui T, Horiuchi S, Endo S, Kakuta R, Ono Y, Yano H.
Data analysis and interpretation: Kakuta N, Nakano R, Nakano A, Suzuki Y, Tanouchi A, Masui T, Horiuchi S, Endo S, Kakuta R, Ono Y, Yano H.
Manuscript writing (original draft): Kakuta N, Nakano R.
Manuscript writing (review and editing): Nakano R, Yano H.
All authors have accepted their responsibility for the entire content of this manuscript and approved submission.
References
Fig. 1
Strategy used for mismatched PCR-RFLP of gyrA and parC QRDRs in A. baumannii. The reverse primers for gyrA and parC are located immediately downstream of the nucleotide sequences corresponding to GyrA87 and ParC84, respectively. The reverse primer for gyrA was designed with one mismatched nucleotide to create an XmnI recognition site (GAANNNNTTC) in the gyrA region containing the codon for Glu-87 (GAA). The reverse primer for parC was designed with two mismatched nucleotides to create an XmnI recognition site in the parC region containing the codon for Glu-84 (GAA). Boldface represents codons 83 and 87 of gyrA and codons 80 and 84 of parC. Underlined DNA sequences indicate restriction sites present in the QRDRs of FQ-susceptible strains.

Fig. 2
PCR-RFLP patterns obtained following digestion with HinfI or XmnI for gyrA and parC. Lanes 1 to 3 and 4 to 6 show PCR-RFLP results for gyrA and parC, respectively. Lane: M, 20 bp DNA ladder marker. (A) PCR-RFLP results for A. baumannii ATCC19606. Lanes: 1, undigested (143 bp); 2, HinfI-digestion (103 bp and 40 bp); 3, XmnI-digestion (122 bp and 21 bp); 4, undigested (120 bp); 5, HinfI-digestion (85 bp and 35 bp); and 6, XmnI-digestion (104 bp and 16 bp). (B) PCR-RFLP results for the representative FQ-resistant A. baumannii strain possessing mutations in gyrA (83) and parC (80). Lanes: 1, undigested (143 bp); 2, HinfI-digestion (143 bp); 3, XmnI-digestion (122 bp and 21 bp); 4, undigested (120 bp); 5, HinfI-digestion (120 bp); and 6, XmnI-digestion (104 bp and 16 bp).
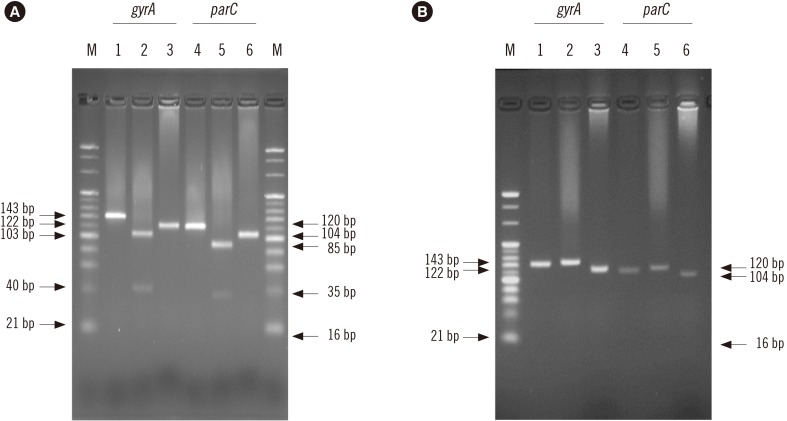
Table 1
Primer sequences and restriction enzymes for mismatched PCR-RFLP
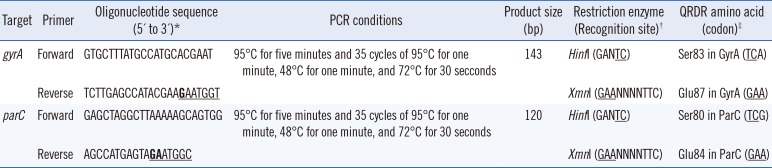
*Boldface represents mismatched nucleotides that introduce artificial restriction sites. Underlined nucleotides indicate restriction sites; †Underlined nucleotides correspond to QRDRs in the gyrA or parC genes; ‡Underlined nucleotides indicate restriction sites.
Abbreviations: QRDR, quinolone resistance-determining region; RFLP, restriction fragment length polymorphism.
Table 2
Levofloxacin and ciprofloxacin MIC ranges, amino acid (codon) changes in GyrA and ParC QRDRs, and mismatched PCR-RFLP results for A. baumannii strains
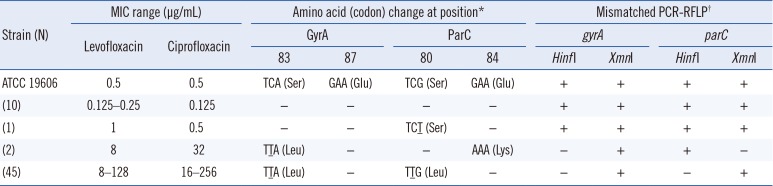
*Compared with ATCC 19606. Underlining indicates point mutations; −, no change.
†+ indicates PCR products that were digested by the restriction enzyme; − indicates PCR products that were not digested by the restriction enzyme.
Abbreviations: QRDR, quinolone resistance-determining region; RFLP, restriction fragment length polymorphism; MIC, minimum inhibitory concentration.




 PDF
PDF ePub
ePub Citation
Citation Print
Print



 XML Download
XML Download