Abstract
Background
Methods
Results
Conclusions
Acknowledgments
References
Fig. 1
Molecular visualization of the wild-type (WT) (left) and variant (right) amino acid using the FoldX function of SNPeffect 4.0. If the mutation destabilizes the structure, ddG (ΔΔG) is increased, whereas stabilizing mutations decrease ddG. Since the FoldX error margin is approximately 0.5 kcal/mol, changes in this range are considered insignificant. (A, B) The mutation from Ile (red in A) to Met (red in B) at position 220 resulted in a ddG of -0.38 kcal/mol. This implies that the mutation had no effect on protein stability. (C, D) The mutation from Gly (red in C) to Ser (red in D) at position 306 resulted in a ddG of 2.81 kcal/mol. This implies that the mutation reduced protein stability. (E, F) The mutation from Ile (red in E) to Thr (red in F) at position 380 resulted in a ddG of 0.56 kcal/mol. This implies that the mutation slightly reduced protein stability. (G, H) The mutation from Cys (red in G) to Gly (red in H) at position 385 resulted in a ddG of 0.65 kcal/mol. This implies that the mutation slightly reduced protein stability. (I, J) The mutation from Arg (red in I) to Cys (red in J) at position 387 resulted in a ddG of 2.18 kcal/mol. This implies that the mutation reduced protein stability. (K, L) The mutation from Pro (red in K) to Arg (red in L) at position 396 resulted in a ddG of 3.19 kcal/mol. This implies that the mutation reduced protein stability. (M and N) The mutation from Glu (red in M) to Lys (red in N) at position 416 resulted in a ddG of -0.86 kcal/mol. This implies that the mutation slightly enhanced protein stability.
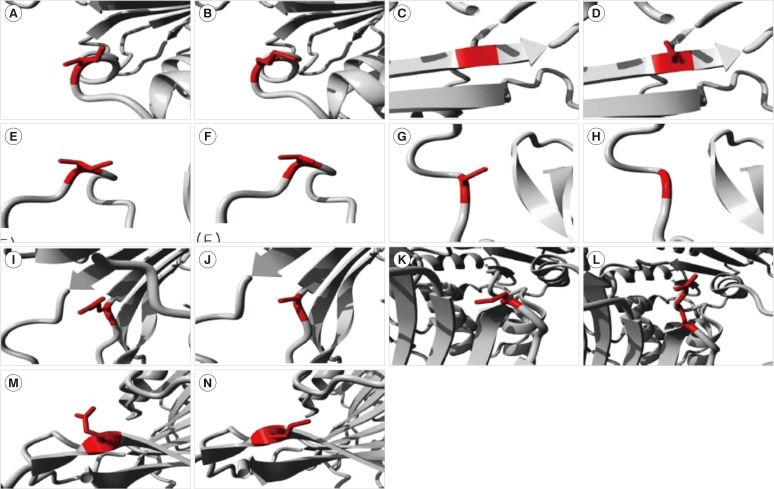
Table 1
Missense mutations in the G6PD reported in Korean patients with G6PD deficiency
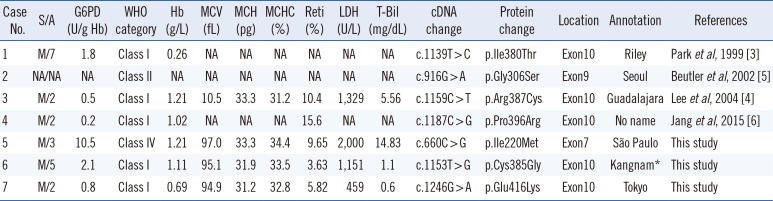
| Case No. | S/A | G6PD (U/g Hb) | WHO category | Hb (g/L) | MCV (fL) | MCH (pg) | MCHC (%) | Reti (%) | LDH (U/L) | T-Bil (mg/dL) | cDNA change | Protein change | Location | Annotation | References |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | M/7 | 1.8 | Class I | 0.26 | NA | NA | NA | NA | NA | NA | c.1139T > C | p.Ile380Thr | Exon10 | Riley | Park et al, 1999 [3] |
| 2 | NA/NA | NA | Class II | NA | NA | NA | NA | NA | NA | NA | c.916G > A | p.Gly306Ser | Exon9 | Seoul | Beutler et al, 2002 [5] |
| 3 | M/2 | 0.5 | Class I | 1.21 | 10.5 | 33.3 | 31.2 | 10.4 | 1,329 | 5.56 | c.1159C > T | p.Arg387Cys | Exon10 | Guadalajara | Lee et al, 2004 [4] |
| 4 | M/2 | 0.2 | Class I | 1.02 | NA | NA | NA | 15.6 | NA | NA | c.1187C > G | p.Pro396Arg | Exon10 | No name | Jang et al, 2015 [6] |
| 5 | M/3 | 10.5 | Class IV | 1.21 | 97.0 | 33.3 | 34.4 | 9.65 | 2,000 | 14.83 | c.660C > G | p.Ile220Met | Exon7 | São Paulo | This study |
| 6 | M/5 | 2.1 | Class I | 1.11 | 95.1 | 31.9 | 33.5 | 3.63 | 1,151 | 1.1 | c.1153T > G | p.Cys385Gly | Exon10 | Kangnam* | This study |
| 7 | M/2 | 0.8 | Class I | 0.69 | 94.9 | 31.2 | 32.8 | 5.82 | 459 | 0.6 | c.1246G > A | p.Glu416Lys | Exon10 | Tokyo | This study |
*Novel mutation identified and annotated in this study.
Abbreviations: G6PD, glucose-6-phosphate dehydrogenase; S/A, sex/age (years); MCH, mean corpuscular hemoglobin; MCV, mean corpuscular volume; MCHC, mean corpuscular hemoglobin concentration; Reti, reticuloscyte; T-Bil, total bilirubin; LDH, lactate dehydrogenase; NA, not available.
Table 2
In-silico analysis of missense mutations of the G6PD gene reported in Korean patients with G6PD deficiency
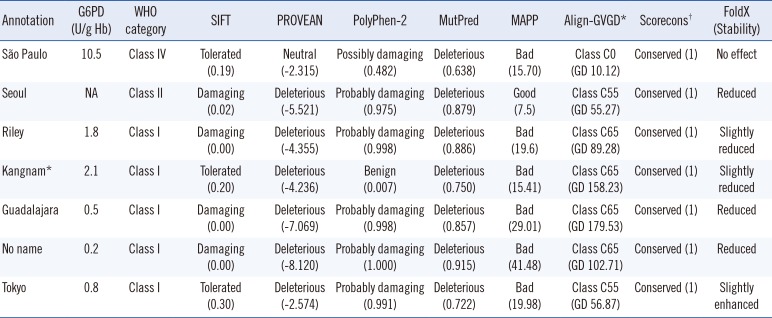
*Novel mutation identified and annotated in this study; †Multiple alignments of different vertebrate species obtained from Evolutionary annotation database (http://www.h-invitational.jp/evola/).
Abbreviations: G6PD, glucose-6-phosphate dehydrogenase; NA, not available.
Table 3
In-silico analysis of missense mutations of G6PD located on same position of amino acid residues identified in Korean G6PD deficiencies
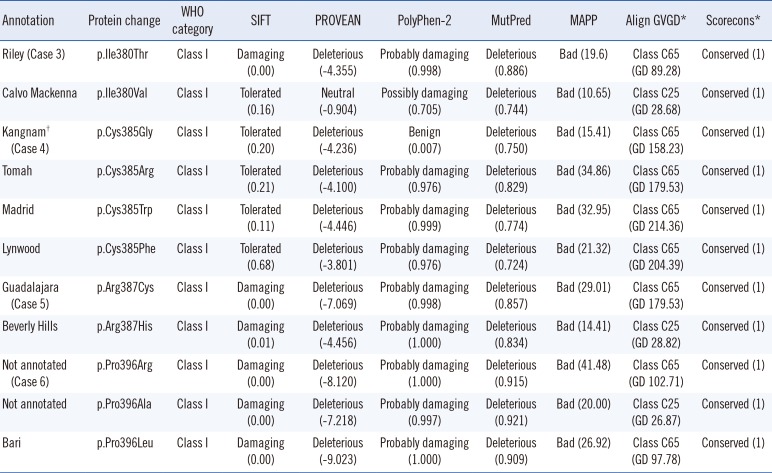
*Multiple alignments of different vertebrate species obtained from Evolutionary annotation database (http://www.h-invitational.jp/evola/); †Novel mutation identified and annotated in this study.
Abbreviation: G6PD, glucose-6-phosphate dehydrogenase.
Table 4
Public variant databases of missense mutations of G6PD located on same position of amino acid residues identified in Korean G6PD deficiencies
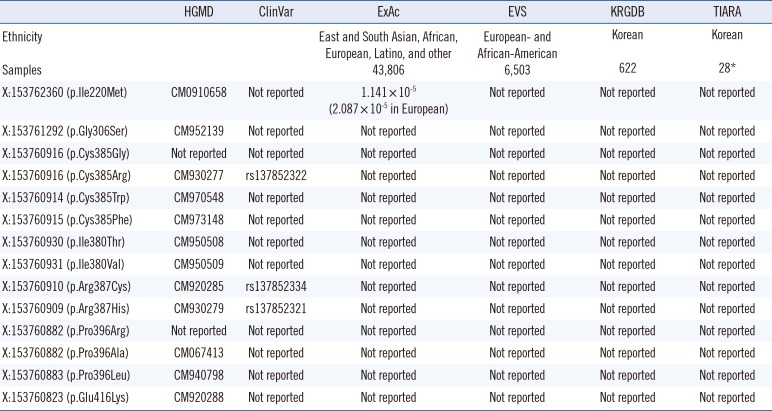
HGMD, Human Gene Mutation Database, http://www.hgmd.cf.ac.uk/.
ClinVar, http://www.ncbi.nlm.nih.gov/clinvar/.
ExAc, The Exome Aggregation Consortium, http://exac.broadinstitute.org/.
EVS, Exome Variant Server, http://evs.gs.washington.edu/.
KRGDB, Korean Reference Genome DB, http://152.99.75.168/KRGDB/.
TIARA, Totally Integrated high-resolution Array cgh data and high-throughput sequencing Read data Archive, http://tiara.gmi.ac.kr/.
*12 whole genomes and 16 transcriptomes
Abbreviation: G6PD, glucose-6-phosphate dehydrogenase.




 PDF
PDF ePub
ePub Citation
Citation Print
Print


 XML Download
XML Download