Abstract
Background
In mammals, the master circadian pacemaker is localized in an area of the ventral hypothalamus known as the suprachiasmatic nucleus (SCN). Previous studies have shown that pacemaker neurons in the SCN are highly coupled to one another, and this coupling is crucial for intrinsic self-sustainability of the SCN central clock, which is distinguished from peripheral oscillators. One plausible mechanism underlying the intercellular communication may involve direct electrical connections mediated by gap junctions.
Methods
We examined the effect of mefloquine, a neuronal gap junction blocker, on circadian Period 2 (Per2) gene oscillation in SCN slice cultures prepared from Per2::luciferase (PER2::LUC) knock-in mice using a real-time bioluminescence measurement system.
The biological clock is a fundamental mechanism coordinating physiological, behavioral, and metabolic processes in a variety of organisms. In mammals, rhythmic biological functions are orchestrated by a circadian pacemaker in the suprachiasmatic nucleus (SCN) of the ventral hypothalamus, which precisely adjusts clock outputs according to solar time through a process of photic synchronization [1234]. Individual neurons within the SCN exhibit independent but highly synchronized circadian oscillations of clock gene expression and neuronal firing [5]. Previous studies suggest that the basic mechanism responsible for the generation of these rhythmic oscillations is an intrinsic property of individual SCN neurons [6].
The core molecular mechanism driving cellular oscillations in the SCN appears to be a feedback loop operating at the transcriptional/translational levels [123]. Although most cells in other extra-SCN brain regions and peripheral tissues also contain their own circadian clock genes with a similar molecular infrastructure, the SCN central clock is required to synchronize circadian rhythms in other tissues. Because of this, it should be noted that the SCN central clock has a distinguishable self-sustainability in its circadian rhythm compared with other peripheral oscillators. Previous studies have shown individual neurons that are dissociated from the SCN have more variable phases and periods, and less reliable rhythms [7], whereas synchrony can be observed at a high density with dispersed neurons. These findings support the hypothesis that the anatomical organization and intercellular communication among SCN neurons may permit the coupling of autonomous cell oscillators to convey coordinated rhythms [56]. Thus, the robust circadian rhythm in the SCN may be attributed to the coupling of SCN neurons forming a stable network of oscillators. However, the detailed mechanisms underlying the circuitry of SCN neurons remain unknown.
The SCN has a heterogeneous structure. Each unilateral SCN consists of about 10,000 neurons in two anatomic subdivisions: the ventrolateral SCN 'core' region, which abuts the optic chiasm and receives retinal input, and the dorsomedial SCN 'shell' region, which partially envelops and receives input from the core [6]. Recently, accumulating evidence suggests that intercellular communication in the SCN is not mediated solely by the synaptic transmission at the synapse, but also through electrical coupling via gap junctions resulting in synchronization of individual cells [8910].
Gap junction channels mediate the intercellular transfer of ions and other small molecules, which allows for electrically and metabolically connected neuronal networks [11]. Gap junction channels are formed by two hemichannels or connexons, which are composed of six connexin proteins. Morphological and electrophysiological studies have indicated that direct communication via gap junctions occurs in SCN neurons [910]. More importantly, mice lacking the connexin (Cx) 36 isoform, which is abundantly expressed in the SCN [12], demonstrate impaired circadian rhythms in their locomotor behaviors [813].
Therefore, we hypothesize that intercellular communication among individual pacemaker neurons is mediated by neuronal gap junctions and by maintenance of clock gene cycling, playing a pivotal role in synchronization. To test this notion, we have employed ex vivo SCN explant cultures based on a knock-in mouse model expressing the Per2::luciferase (PER2::LUC) fusion protein. We examined the effect of mefloquine, a neuronal gap junction inhibitor [14], on the circadian oscillation of PER2::LUC at the levels of both whole SCN and individual neurons.
Adult mPer2::luciferase knock-in mice [15] were maintained in temperature-controlled (22℃ to 23℃) quarters under a 12 hours light and 12 hours dark (light-dark) photoperiod (light-on at 8:00 AM) with standard mouse chow and water available ad libitum. Heterozygous or homozygous male adult mice were mated with wild-type females to yield PER2::LUC-expressing pups. All animal procedures were approved by the Institutional Animal Care and Use Committee of Seoul National University.
Mefloquine (Sigma, St. Louis, MO, USA) was dissolved in dimethyl sulfoxide (DMSO). Mefloquine was directly added to culture medium after the 3rd or 4th peak appeared during experiments.
The slice culture was prepared as previously reported with minor modifications [16]. Neonatal (5- to 10-day-old) PER2::LUC mice were sacrificed, and the brains were quickly removed. The brain was coronally sectioned at a thickness of 400 µm with a vibratome (Campden Instruments, Leicester, UK). The slices were maintained on a culture insert membrane (Millicell-CM, Millipore, Bedford, MA, USA) and dipped into culture medium (50% minimum essential medium, 25% Gey's balanced salt solution, 25% horse serum, 36 mmol/L glucose, and 100 units/mL aerosolized antibiotics) at 37℃. The SCN slices were cultivated for 2 weeks before being employed in experiments. Peripheral tissues (liver, kidney, lung, heart, and adrenal glands) were sectioned at 1 mm thickness and used as control preparations.
The bioluminescence from the SCN slice cultures was monitored as previously reported with slight modifications [17]. The SCN slice cultures from the PER2::LUC mice were maintained in sealed 35 mm petri dishes with 1 mL of culture medium containing 0.3 mmol/L D-luciferin (Promega, Madison, WI, USA) at 36℃. The light emission was measured and integrated for 2 minutes at 20 minutes intervals with a dish-type wheeled luminometer (AB-2550 Kronos-Dio, ATTO, Tokyo, Japan). For recording bioluminescence from peripheral tissues, the tissues were synchronized with a brief application of dexamethasone (DEX; 1 µM) for 2 hours before beginning bioluminescence recordings (Fig. 1).
Bioluminescence imaging of individual cells in brain slice cultures was slightly modified, as previously reported [16]. The SCN slice cultures from PER2::LUC mice were maintained in a customized teflon chamber (Live Cell Instrument, Seoul, Korea) with two input ports and one output port. Culture medium containing 0.3 mmol/L D-luciferin was perifused at a flow rate of 0.4 mL/hr. For measuring the bioluminescence from SCN slice cultures at the single cell level, we used an imaging system with an ultrasensitive charge-coupled device (CCD) camera coupled to a perifusion system for continuous media supply (Cellgraph AB-1000, ATTO, Tokyo, Japan). An IBM personal computer interfaced to a two dimensional photon-counting camera continuously recorded bioluminescence data for 10 minutes.
Period and phase measurements were calculated as described in previous reports [1718]. Briefly, data (2-minute bins) were smoothed by an adjacent-averaging method with 1.67 hours running means. The peak was calculated as the highest point of the smoothed data using the CLUSTER8 program (Veldhuis, University of Virginia, Charlottesville, VA, USA), and this statistical analysis program identifies significant interval values for peak and nadir. Peaks and amplitudes were normalized by dividing the difference of a peak and nearby nadir by the average intensity of the basal section. Period stability was examined using the standard deviation of mean periods from each experiment. For the raster plots, normalized bioluminescence intensity data were normalized by standard deviation and were color-coded in red for a standard deviation higher than 1.5 and green for a standard deviation lower than 1.5. Plots were constructed with the Hierarchical Clustering Explorer 3.0 (HCE3) program.
To determine the endogenous circadian characteristics of organotypic cultures from various tissues, PER2::LUC expression was measured with a quantitative real-time bioluminescence monitoring system. The PER2::LUC knock-in mice have a firefly luciferase gene fused in-frame to the 3' end of the endogenous Per2 gene [15]. Neonatal PER2::LUC knock-in mice were used to prepare the SCN explant cultures. The SCN culture displayed robust and sustained circadian oscillation of PER2::LUC expression for up to eight days with a mean period of 24.56±0.12 hours (Fig. 1A). Circadian oscillations of PER2::LUC expression were also examined in several peripheral tissues, including liver, kidney, lung, heart, pituitary, and adrenal glands (Fig. 1B). For synchronization of local oscillators, all peripheral tissues were treated with DEX (1 µM) 2 hours before bioluminescence recording. A gradual decrease in the circadian amplitudes was observed because of the depletion of both nutrients and D-luciferin substrates. However, compared with the SCN explant, PER2::LUC oscillatory profiles in peripheral tissues exhibited relatively lower amplitudes and tended to more rapidly attenuate as time went on, presumably due to the out-of-phase individual cellular rhythms.
To examine whether intercellular communication via gap junctions may underlie the robustness and/or sustainability of periodic clock gene expression in whole SCN tissue, we examined the effect of chronic treatment with the neuronal gap junction inhibitor mefloquine on PER2::LUC expression in SCN cultures (Fig. 2). Treatment with vehicle (0.1% DMSO) for 4 days evoked no obvious changes in the phase and period of the rhythm. Changing medium did appear to increase overall bioluminescence signals, but this was likely due to fresh substrate supply. Interestingly, mefloquine application during a trough in Per2 promoter-driven luciferase activity elicited phase-advancing effects (0.32±0.18 hours for controls, n=8; 1.79±0.67 hours for 3 µM and 3.08±0.36 hours for 10 µM mefloquine, n=6 for each dose). However, apparent reductions in the amplitude of the rhythm, as well as unstable rhythmicity were seen in the presence of 10 µM mefloquine.
To further define the effect of mefloquine, the rhythmic features were statistically evaluated, including periods, stability of the rhythm, and amplitudes during pre- (before treatment), treat- (during treatment), and post- (after washout) periods of the treatment. The mean periods of the rhythm, as determined by averaging peak-to-peak intervals in a set of experiments except the transitional phases, showed no significant differences at approximately 24.5 hours in all conditions (Fig. 3A). However, treatment with 10 µM mefloquine impaired the period stability, as determined by the variations in peak-to-peak intervals in a set of experiments, which showed less coherent rhythmicities than those observed in the control and 3 µM mefloquine groups (Fig. 3B). The standard deviation of the periods of an explant increased to ±2.05 hours during the treatment with 10 µM mefloquine. In addition, chronic treatment with mefloquine at a higher dose reduced the amplitude of the oscillating bioluminescence. The amplitudes decreased to 54.17%±1.72% of the pretreatment period (mean±% SEM, n=6; the mean amplitude during the pretreatment period was set at 100%) in the presence of 10 µM mefloquine (Fig. 3C). This is a significantly larger reduction in amplitude compared with the control and 3 µM mefloquine groups (74.12%±5.31% for control, n=8; 82.59%±5.39% for 3 µM mefloquine treatment, n=6). These findings clearly indicate that inhibition of neuronal gap junctions in the SCN weakens the circadian rhythm of PER2::LUC expression. Furthermore, the effect of mefloquine is unlikely a result of treatment toxicity, because the weakened rhythmicity of PER2::LUC expression was restored by drug-washout.
In the next set of experiments, we examined whether the impaired circadian PER2::LUC expression in the whole SCN explant is due to weakened synchrony of individual cellular rhythms. For this purpose, bioluminescence signals from the SCN explant were monitored at a single cell level using another bioluminescence real-time imaging system (Fig. 4). We examined the effect of chronic treatment with 10 µM mefloquine on circadian oscillation of PER2::LUC expression at the whole SCN level. Similar to the results obtained with the quantitative bioluminescence recording method, the periodicity tended to be unstable after mefloquine application (Fig. 4). The bioluminescence signal intensities and the amplitudes of the rhythm were also decreased after treatment, but the spontaneous decline in signal intensities and amplitudes in the static incubation system was less apparent due to the continuous supply of fresh recording medium during live cell imaging.
Furthermore, the individual cellular rhythms and their synchrony were analyzed from 80 randomly chosen cells throughout the SCN, and the bioluminescence signals from each cell were plotted (Fig. 5A). For most cells, mefloquine administration reduced the amplitude of the rhythms by lowering the peak bioluminescence intensity in each cycle, although the individual mean periods were not significantly altered (Fig. 5B). Impaired stability of the rhythm was also demonstrated at the single cell level, as expressed by variations in the peak-to-peak intervals. More importantly, intercellular synchrony in the SCN was apparently impaired by the inhibition of gap junctions (Fig. 5B-D). A raster plot presentation of bioluminescence signals showed more variable locations of peaks and troughs among individual cells during mefloquine administration (Fig. 5C). Phase differences among individual cells compared with the peak time at the whole SCN level also supports the impairment in synchrony (Fig. 5D). Collectively, these results demonstrate that inhibition of neuronal gap junctions or electrical synapses in the SCN results in attenuated and unstable circadian cycles in the molecular clockwork of individual pacemaker neurons as well as their overall synchrony, thereby weakening the periodicity of the central clock.
The present study showed that neuronal gap junctions play an important role in regulating circadian rhythms of the SCN central clock by influencing cyclic clock gene expression in individual cells as well as the synchrony of the individual rhythms. The involvement of electrical synapses for controlling circadian function of the hypothalamic SCN has been previously proposed based on neuropeptide production and behavioral rhythms [8910]. The SCN is composed of networks formed by thousands of pacemaker neurons, which can act individually as autonomous oscillators. These multiple oscillators in the SCN normally function in a unified manner. Growing evidence indicates that connectivity among SCN neurons is not only essential for proper time-keeping, but also confers stability and robustness of the SCN as a central pacemaker for the circadian timing system [6]. In this regard, it should be noted that SCN central pacemaker neurons and other peripheral cells contain molecular circadian oscillators with a similar molecular makeup [123]. In fact, it is of considerable interest how the SCN can generate more autonomous and self-sustainable rhythms. Several ion channels, such as voltage-gated Na+ and L-type Ca2+ channels, as well as paracrine neuropeptides, including vasoactive intestinal polypeptide and arginine vasopressin, have been proposed to confer the exceptional stability of the SCN central clock [10192021].
It has been recently shown that about 25% of the SCN neurons are electrically coupled and show synchronized spiking activities [810]. Several other studies suggested the role of glial gap junctions in the SCN [11]. In the SCN, Cx36 may be a primary subunit in neural gap junctions as revealed by immunohistochemical and electromicroscopic studies, while Cx30, Cx32, and Cx43 have been observed in glial cells [12]. In contrast to general gap junction inhibitors acting on a broad range of connexin subunits, mefloquine inhibits only Cx36 and Cx50-containing channels, which are enriched in neuronal cells. When expressed in transfected N2A neuroblastoma cells, mefloquine efficiently blocked Cx36 and Cx50-containing channels even at very low concentrations (IC50=1.1 µM), although it can inhibit other gap junction subunits, including Cx43, Cx32, and Cx26, at 10- to 100-fold higher concentrations [14].
As shown in Figs. 2, 3, the instability and reduced robustness of circadian gene cycling are the most characteristic consequences of chronic mefloquine treatment. Increased variations in the periods but reduced amplitudes of cyclic PER2::LUC expression are found at both the whole SCN and single cell levels. Weakened synchrony among individual cells is also likely to underlie impaired rhythm stability. These findings are consistent with previous studies showing that pharmacological or genetic abrogation of the functional gap junctions in the SCN can impair stability as well as the amplitude of the circadian locomotor rhythms [811]. Because Cx36 is known as the major component of neuronal electrical coupling and also mediates abnormal behavior rhythms [8], the most probable molecular target of mefloquine for altering the PER2 rhythm is Cx36-containing connexon hemichannels. It is noteworthy that Cx43, Cx32, and Cx26 are also known as targets of mefloquine. Interestingly, these three subunits of connexin are mainly expressed in peripheral tissues such as heart (Cx43) and liver (Cx32, Cx26) [22] unlike to Cx36. But the affinity of mefloquine to these three subunits are 10 to 100 times lower than to Cx36 [14]. Also, circadian rhythm in peripheral tissues is much less stable and is basically dependent on presence of the SCN [1]. Although importance of gap junctions in peripheral clocks still remains elusive, these low stability and dependency to central clock suggest that electrical connectivity between cells in peripheral tissues may have limited roles in regulation of circadian rhythm as compared to that in the SCN. Therefore, a low concentration of mefloquine (about 10 µM) would not cause to inhibit gab junction activity or to alter circadian rhythm in peripheral tissues but only would be effective in central nervous system including the SCN. This possibility should be further investigated with additional subunit-specific abrogation experiments.
The present study examined the effects of mefloquine on synchronized circadian clock gene oscillations in the SCN. However, its putative responsiveness to clock-resetting signals also requires further testing. Although the phase of the SCN central rhythm can be reset by certain stimuli, such as glutamate, serotonin, and cAMP [17232425], it is quite resistant to other signals, such as glucocorticoid and temperature pulse [2627]. In the physiological context, such resistance helps organisms stabilize their intrinsic rhythmicity against sporadic environmental cues [28]. Takahashi and colleagues have demonstrated that activity-dependent ion channels and stochastic mechanisms based on neural connectivity are heavily involved in the unusual resistance of the central rhythm generated by the SCN [2729]. Therefore, it should be determined whether intercellular connectivity by neuronal gap junctions also contributes to the robustness of the SCN central rhythm.
In summary, the present study provides experimental evidence for the role of neuronal population coupling via neuronal gap junctions for harmonizing the robust and stable clock gene cycling in the SCN. Further studies are needed to dissect the anatomical localization of connexin subunits within the intra-SCN neural circuits to gain insight into their functional relevance for the circadian timing system.
Figures and Tables
Fig. 1
Real-time analyses of circadian Per2::luciferase (PER2::LUC) expression in suprachiasmatic nucleus (SCN) and peripheral tissues explants ex vivo. (A) A representative circadian profile of PER2::LUC expression in the SCN explant culture. Daily periods were calculated and are expressed during 8-day and cultivation periods at the top. (B) Circadian profiles of PER2::LUC expression in peripheral tissues ex vivo. Results from the liver, kidney, lung, heart, pituitary, and adrenal glands are shown. Peripheral tissues were synchronized by a brief application of dexamethasone (DEX, 1 µM) for 2 hours (arrows).
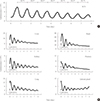

Fig. 2
Chronic treatment with mefloquine attenuates the circadian rhythm of Per2::luciferase (PER2::LUC) expression in the suprachiasmatic nucleus. Representative bioluminescence recordings are shown in (A) vehicle (0.1% dimethyl sulfoxide), (B) 3 and (C) 10 µM mefloquine-treated groups. Mefloquine was applied directly into the recording medium at a trough after the 3rd or 4th peak. After four more cycles passed, the medium was changed. Daily periods are calculated and expressed during 11- or 12-day cultivation periods at the top of each figure.

Fig. 3
Effect of mefloquine on Per2::luciferase (PER2::LUC) cycling activities in the suprachiasmatic nucleus. Mean periods (A), period stability (B, SD of mean periods in a cell), and changes in amplitude (C, % of pretreatment) were statistically analyzed among pre- (before treatment), treat- (during treatment), and post- (after washout) periods in three groups (vehicle [n=8], 3 µM [n=6], and 10 µM [n=6] mefloquine) on the rhythmic gene expression of Per2. Data are expressed as mean±standard error of the mean. aP<0.01 vs. pretreatment.
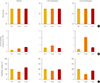
Fig. 4
Bioluminescence-based real-time imaging of Per2::luciferase (PER2::LUC) reporter activity in the presence of mefloquine. (A) Representative image of the PER2::LUC-expressing suprachiasmatic nucleus (SCN) taken with an intensified CCD camera in the presence of perfused recording medium containing 0.3 mmol/L D-luciferin. (B) Bioluminescence traces measured from a whole PER2::LUC SCN culture. After the 3rd peak of the rhythm, 10 µM mefloquine was applied during the trough. The black bar indicates the period of chronic treatment with 10 µM mefloquine. (C) Time-lapse images show circadian fluctuations of PER2::LUC expression in the SCN culture. Each image was taken by integrating the bioluminescence for 10 minutes. The images were aligned from every six images (every hour) for 7 days.
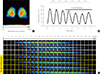

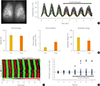
Fig. 5
Attenuated circadian Per2::luciferase (PER2::LUC) expression and synchrony among individual suprachiasmatic nucleus (SCN) cells. (A) The video-micrograph illustrates 80 randomly chosen individual cells in the SCN (left panel). The right panel displays the bioluminescence recordings from individual SCN cells. The black bar indicates the chronic treatment with 10 µM mefloquine. (B) Bar charts summarizing the effects of mefloquine at the single-cell level. Mean periods among examined cells, period stability (SD of mean periods in a cell), and changes in amplitude (% of pretreatment) are expressed as the mean±standard error of the mean. (C) A raster plot presentation displaying voxels measured dorsal to ventral throughout the SCN. Red corresponds to the peak of bioluminescence and green to the trough. (D) Phase differences in peak time of each cell compared to the mean value. The X axis represents the order of peaks (seven total peaks are shown). The Y axis represents differences in peak time (hour) of individual cells, shown dotted. SD indicates index variations. aP<0.01 vs. pretreatment.
ACKNOWLEDGMENTS
This work was supported by grants from the Korea Ministry of Education, Science, and Technology (MEST) through the Brain Research Center of the 21st Century Frontier Research Program (2009K001287), the Brain Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (NRF-2013M3C7A 1056731) and the BK21 Plus program through the National Research Foundation of Korea funded by the Ministry of Education (10Z20130012420).
References
1. Dibner C, Schibler U, Albrecht U. The mammalian circadian timing system: organization and coordination of central and peripheral clocks. Annu Rev Physiol. 2010; 72:517–549.
2. Son GH, Chung S, Kim K. Biological rhythms and neuroendocrine systems. Endocrinol Metab (Seoul). 2010; 25:249–257.
3. Mohawk JA, Green CB, Takahashi JS. Central and peripheral circadian clocks in mammals. Annu Rev Neurosci. 2012; 35:445–462.
4. Girardet C, Becquet D, Blanchard MP, Francois-Bellan AM, Bosler O. Neuroglial and synaptic rearrangements associated with photic entrainment of the circadian clock in the suprachiasmatic nucleus. Eur J Neurosci. 2010; 32:2133–2142.
5. Colwell CS. Rhythmic coupling among cells in the suprachiasmatic nucleus. J Neurobiol. 2000; 43:379–388.
6. Welsh DK, Takahashi JS, Kay SA. Suprachiasmatic nucleus: cell autonomy and network properties. Annu Rev Physiol. 2010; 72:551–577.
7. Herzog ED, Aton SJ, Numano R, Sakaki Y, Tei H. Temporal precision in the mammalian circadian system: a reliable clock from less reliable neurons. J Biol Rhythms. 2004; 19:35–46.
8. Long MA, Jutras MJ, Connors BW, Burwell RD. Electrical synapses coordinate activity in the suprachiasmatic nucleus. Nat Neurosci. 2005; 8:61–66.
9. Shinohara K, Funabashi T, Mitushima D, Kimura F. Effects of gap junction blocker on vasopressin and vasoactive intestinal polypeptide rhythms in the rat suprachiasmatic nucleus in vitro. Neurosci Res. 2000; 38:43–47.
10. Wang MH, Chen N, Wang JH. The coupling features of electrical synapses modulate neuronal synchrony in hypothalamic superachiasmatic nucleus. Brain Res. 2014; 1550:9–17.
11. McCracken CB, Roberts DC. Neuronal gap junctions: expression, function, and implications for behavior. Int Rev Neurobiol. 2006; 73:125–151.
12. Rash JE, Olson CO, Pouliot WA, Davidson KG, Yasumura T, Furman CS, et al. Connexin36 vs. connexin32, "miniature" neuronal gap junctions, and limited electrotonic coupling in rodent suprachiasmatic nucleus. Neuroscience. 2007; 149:350–371.
13. Juszczak GR, Swiergiel AH. Properties of gap junction blockers and their behavioural, cognitive and electrophysiological effects: animal and human studies. Prog Neuropsychopharmacol Biol Psychiatry. 2009; 33:181–198.
14. Cruikshank SJ, Hopperstad M, Younger M, Connors BW, Spray DC, Srinivas M. Potent block of Cx36 and Cx50 gap junction channels by mefloquine. Proc Natl Acad Sci U S A. 2004; 101:12364–12369.
15. Yoo SH, Yamazaki S, Lowrey PL, Shimomura K, Ko CH, Buhr ED, et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc Natl Acad Sci U S A. 2004; 101:5339–5346.
16. Choe HK, Kim HD, Park SH, Lee HW, Park JY, Seong JY, et al. Synchronous activation of gonadotropin-releasing hormone gene transcription and secretion by pulsatile kisspeptin stimulation. Proc Natl Acad Sci U S A. 2013; 110:5677–5682.
17. Asai M, Yamaguchi S, Isejima H, Jonouchi M, Moriya T, Shibata S, et al. Visualization of mPer1 transcription in vitro: NMDA induces a rapid phase shift of mPer1 gene in cultured SCN. Curr Biol. 2001; 11:1524–1527.
18. Abe M, Herzog ED, Yamazaki S, Straume M, Tei H, Sakaki Y, et al. Circadian rhythms in isolated brain regions. J Neurosci. 2002; 22:350–356.
19. Aton SJ, Colwell CS, Harmar AJ, Waschek J, Herzog ED. Vasoactive intestinal polypeptide mediates circadian rhythmicity and synchrony in mammalian clock neurons. Nat Neurosci. 2005; 8:476–483.
20. Irwin RP, Allen CN. Neuropeptide-mediated calcium signaling in the suprachiasmatic nucleus network. Eur J Neurosci. 2010; 32:1497–1506.
21. Maywood ES, Chesham JE, O'Brien JA, Hastings MH. A diversity of paracrine signals sustains molecular circadian cycling in suprachiasmatic nucleus circuits. Proc Natl Acad Sci U S A. 2011; 108:14306–14311.
22. Sohl G, Willecke K. An update on connexin genes and their nomenclature in mouse and man. Cell Commun Adhes. 2003; 10:173–180.
23. Mizoro Y, Yamaguchi Y, Kitazawa R, Yamada H, Matsuo M, Fustin JM, et al. Activation of AMPA receptors in the suprachiasmatic nucleus phase-shifts the mouse circadian clock in vivo and in vitro. PLoS One. 2010; 5:e10951.
24. O'Neill JS, Maywood ES, Chesham JE, Takahashi JS, Hastings MH. cAMP-dependent signaling as a core component of the mammalian circadian pacemaker. Science. 2008; 320:949–953.
25. Ciarleglio CM, Resuehr HE, McMahon DG. Interactions of the serotonin and circadian systems: nature and nurture in rhythms and blues. Neuroscience. 2011; 197:8–16.
26. Balsalobre A, Brown SA, Marcacci L, Tronche F, Kellendonk C, Reichardt HM, et al. Resetting of circadian time in peripheral tissues by glucocorticoid signaling. Science. 2000; 289:2344–2347.
27. Buhr ED, Yoo SH, Takahashi JS. Temperature as a universal resetting cue for mammalian circadian oscillators. Science. 2010; 330:379–385.
28. Kiessling S, Eichele G, Oster H. Adrenal glucocorticoids have a key role in circadian resynchronization in a mouse model of jet lag. J Clin Invest. 2010; 120:2600–2609.
29. Ko CH, Yamada YR, Welsh DK, Buhr ED, Liu AC, Zhang EE, et al. Emergence of noise-induced oscillations in the central circadian pacemaker. PLoS Biol. 2010; 8:e1000513.




 PDF
PDF ePub
ePub Citation
Citation Print
Print


 XML Download
XML Download