Abstract
The rotavirus nonstructural glycoprotein, NSP4, has been identified as the first viral enterotoxin capable of inducing diarrhea. To investigate the biological function of NSP4 in the inflammatory process, a cDNA from human rotavirus (Wa strain) RNA segment 10 was amplified by RT-PCR, cloned into TA vector, and subsequently subcloned into pET23b expression plasmid. The expression of NSP4 protein was determined by SDS-PAGE and Western blotting, then, the protein was purified by affinity chromatography on Ni-NTA-agarose column. The inflammatory effects of NSP4, namely, production of nitric oxide (NO), proinflammatory cytokines (IL-1β, IL-6, IL-10, and TNF-α), and prostaglandin E2 (PGE2), was evaluated using NSP4-stimulated RAW 264.7 murine macrophages and compared with those observed after stimulation with lipopolysaccharide (LPS). The levels of IL-1β, IL-6, and TNF-α were significantly increased, and those of NO and PGE2 also increased in NSP4-stimulated RAW 264.7 cells. These findings indicate that NSP4 plays an important role in the inflammatory response observed during rotavirus infection.
Go to : 
REFERENCES
1). Kapikian AZ, Hoshino Y, Chanock RM. Rotaviruses. Knipe D, Howley P, Griffin D, Lamb R, Martin M, Straus S, editors. Fields Virology. 4th ed.Philadelphia: Lippincott, Williams & Wilkins;2001. p. 1787–833.
2). Parashar UD, Burton A, Lanata C, Boschi-Pinto C, Shibuya K, Steele D, et al. Global mortality associated with rotavirus disease among children in 2004. J Infect Dis. 2009; 200(Suppl 1):S9–S15.

3). Hertel PM, Estes MK. Rotavirus and biliary atresia: can causation be proven? Curr Opin Gastroenterol. 2012; 28:10–7.
4). Ball JM, Tian P, Zeng CQ, Morris AP, Estes MK. Age-dependent diarrhea induced by a rotaviral nonstructural glycoprotein. Science. 1996; 272:101–4.

5). Dong Y, Zeng CQ, Ball JM, Estes MK, Morris AP. The rotavirus enterotoxin NSP4 mobilizes intracellular calcium in human intestinal cells by stimulating phospholipase C-mediated inositol 1, 4, 5-trisphosphate production. Proc Natl Acad Sci U S A. 1997; 94:3960–5.
6). Díaz Y, Peña F, Aristimuño OC, Matteo L, De Agrela M, Chemello ME, et al. Dissecting the Ca(2)(+) entry pathways induced by rotavirus infection and NSP4-EGFP expression in Cos-7 cells. Virus Res. 2012; 167:285–96.

7). Rodríguez-Diaz J, López-Andújar P, García-Díaz A, Cuenca J, Montava R, Buesa J. Expression and purification of polyhistidine-tagged rotavirus NSP4 proteins in insect cells. Protein Expr Purif. 2003; 31:207–12.
8). Díaz Y, Chemello ME, Peña F, Aristimuño OC, Zambrano JL, Rojas H, et al. Expression of nonstructural rotavirus protein NSP4 mimics Ca2+ homeostasis changes induced by rotavirus infection in cultured cells. J Virol. 2008; 82:11331–43.
9). Warfield KL, Blutt SE, Crawford SE, Kang G, Conner ME. Rotavirus infection enhances lipopolysaccharide-induced intussusception in a mouse model. J Virol. 2006; 80:12377–86.

10). Ware CF. Network communications: lymphotoxins, LIGHT, and TNF. Annu Rev Immunol. 2005; 23:787–819.

11). Ulevitch RJ, Tobias PS. Receptor-dependent mechanisms of cell stimulation by bacterial endotoxin. Annu Rev Immunol. 1995; 13:437–57.

12). Harris SG, Padilla J, Koumas L, Ray D, Phipps RP. Prostaglandins as modulators of immunity. Trends Immunol. 2002; 23:144–50.

13). Boveris A, Alvarez S, Navarro A. The role of mitochondrial nitric oxide synthase in inflammation and septic shock. Free Radic Biol Med. 2002; 33:1186–93.

14). Borghan MA, Mori Y, El-Mahmoudy AB, Ito N, Sugiyama M, Takewaki T, et al. Induction of nitric oxide synthase by rotavirus enterotoxin NSP4: implication for rotavirus pathogenicity. J Gen Virol. 2007; 88:2064–72.

15). Ge Y, Mansell A, Ussher JE, Brooks AE, Manning K, Wang CJ, et al. Rotavirus NSP4 Triggers Secretion of Proinflammatory Cytokines from Macrophages via Toll-Like Receptor 2. J Virol. 2013; 87:11160–7.

16). Browne EP, Bellamy AR, Taylor JA. Membrane-destabilizing activity of rotavirus NSP4 is mediated by a membrane-proximal amphipathic domain. J Gen Virol. 2000; 81:1955–9.

17). Sharifi Z, Yakhchali B, Shahrabadi MS. Expression and one step purification of the full-length biologically active, NSP4 of human rotavirus Wa strain. Int J Mol Med Adv Sci. 2005; 1:206–12.
18). Tabandeh F, Sanati MH, Shoja Alsadati SA, Yakhchali B, Khodabandeh M. Evaluation of heat induction strategy for recombinant human growth hormone expression in fed-batch fermentation. Iran J Biotechnology. 2005; 3:24–30.
19). Mori Y, Borgan MA, Ito N, Sugiyama M, Minamoto N. Sequential analysis of nonstructural protein NSP4s derived from Group A avian rotaviruses. Virus Res. 2002; 89:145–51.

20). Johansen K, Hinkula J, Espinoza F, Levi M, Zeng C, Rudén U, et al. Humoral and cell-mediated immune responses in humans to the NSP4 enterotoxin of rotavirus. J Med Virol. 1999; 59:369–77.

21). Iosef C, Chang KO, Azevedo MS, Saif LJ. Systemic and intestinal antibody responses to NSP4 enterotoxin of Wa human rotavirus in a gnotobiotic pig model of human rotavirus disease. J Med Virol. 2002; 68:119–28.

22). Rodríguez-Díaz J, Banasaz M, Istrate C, Buesa J, Lundgren O, Espinoza F, et al. Role of nitric oxide during rotavirus infection. J Med Virol. 2006; 78:979–85.

23). Allen SR, Jafri M, Donnelly B, McNeal M, Witte D, Bezerra J, et al. Effect of rotavirus strain on the murine model of biliary atresia. J Virol. 2007; 81:1671–9.

24). Rauschenfels S, Krassmann M, Al-Masri AN, Verhagen W, Leonhardt J, Kuebler JF, et al. Incidence of hepatotropic viruses in biliary atresia. Eur J Pediatr. 2009; 168:469–76.

25). Hertel PM, Crawford SE, Finegold MJ, Estes MK. Osteopontin upregulation in rotavirus-induced murine biliary atresia requires replicating virus but is not necessary for development of biliary atresia. Virology. 2011; 417:281–92.

26). Wang W, Donnelly B, Bondoc A, Mohanty SK, McNeal M, Ward R, et al. The rhesus rotavirus gene encoding VP4 is a major determinant in the pathogenesis of biliary atresia in newborn mice. J Virol. 2011; 85:9069–77.

27). Feng J, Yang J, Zheng S, Qiu Y, Chai C. Silencing of the rotavirus NSP4 protein decreases the incidence of biliary atresia in murine model. PLoS One. 2011; 6:e23655.

28). Yang H, Plösch T, Lisman T, Gouw AS, Porte RJ, Verkade HJ, et al. Inflammation mediated down-regulation of hepatobiliary transporters contributes to intrahepatic cholestasis and liver damage in murine biliary atresia. Pediatr Res. 2009; 66:380–5.

29). Vernooy JH, Dentener MA, van Suylen RJ, Buurman WA, Wouters EF. Long-term intratracheal lipopolysaccharide exposure in mice results in chronic lung inflammation and persistent pathology. Am J Respir Cell Mol Biol. 2002; 26:152–9.

30). Bondeson J. The mechanisms of action of disease-modifying antirheumatic drugs: a review with emphasis on macrophage signal transduction and the induction of proinflammatory cytokines. Gen Pharmacol. 1997; 29:127–50.

31). Kinne RW, Bräuer R, Stuhlmüller B, Palombo-Kinne E, Burmester GR. Macrophages in rheumatoid arthritis. Arthritis Res. 2000; 2:189–202.
32). Dayer JM. The process of identifying and understanding cytokines: from basic studies to treating rheumatic diseases. Best Pract Res Clin Rheumatol. 2004; 18:31–45.

33). Diehl S, Rincón M. The two faces of IL-6 on Th1/Th2 differentiation. Mol Immunol. 2002; 39:531–6.

34). Scheller J, Chalaris A, Schmidt-Arras D, Rose-John S. The proand anti-inflammatory properties of the cytokine interleukin-6. Biochim Biophys Acta. 2011; 1813:878–88.
Go to : 
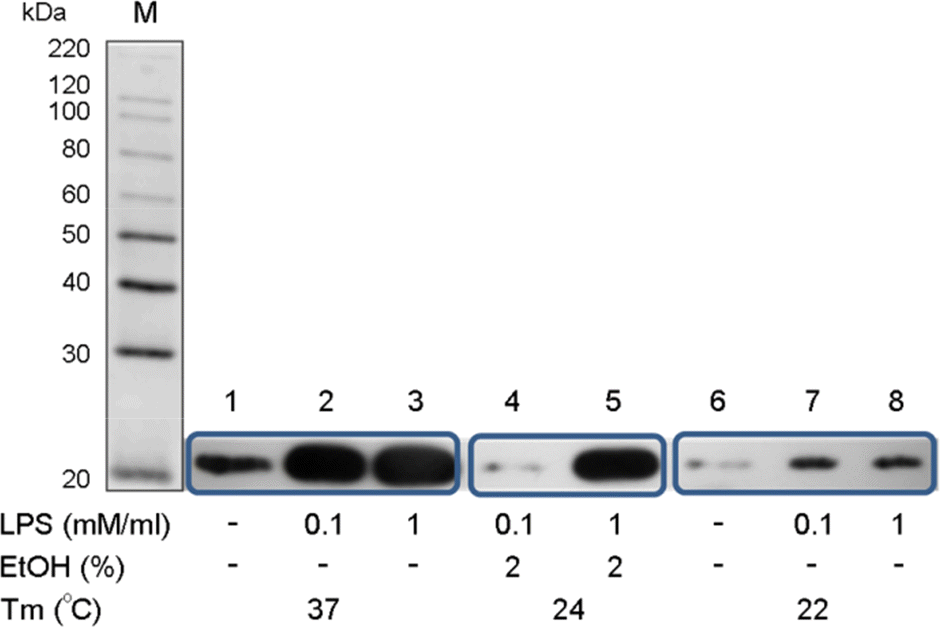 | Figure 1.Expression of recombinant NSP4. Recombinant NSP4 was expressed by induction with IPTG, as shown by western blotting: lane 1~3: 0, 0.1, and 1 mM IPTG at 37°C, respectively; lane 4~5: 0 and 1 mM IPTG at 37°C, 2% EtOH; and lane 6~8: 0, 0.1, and 1 mM IPTG at 22°C, respectively. |
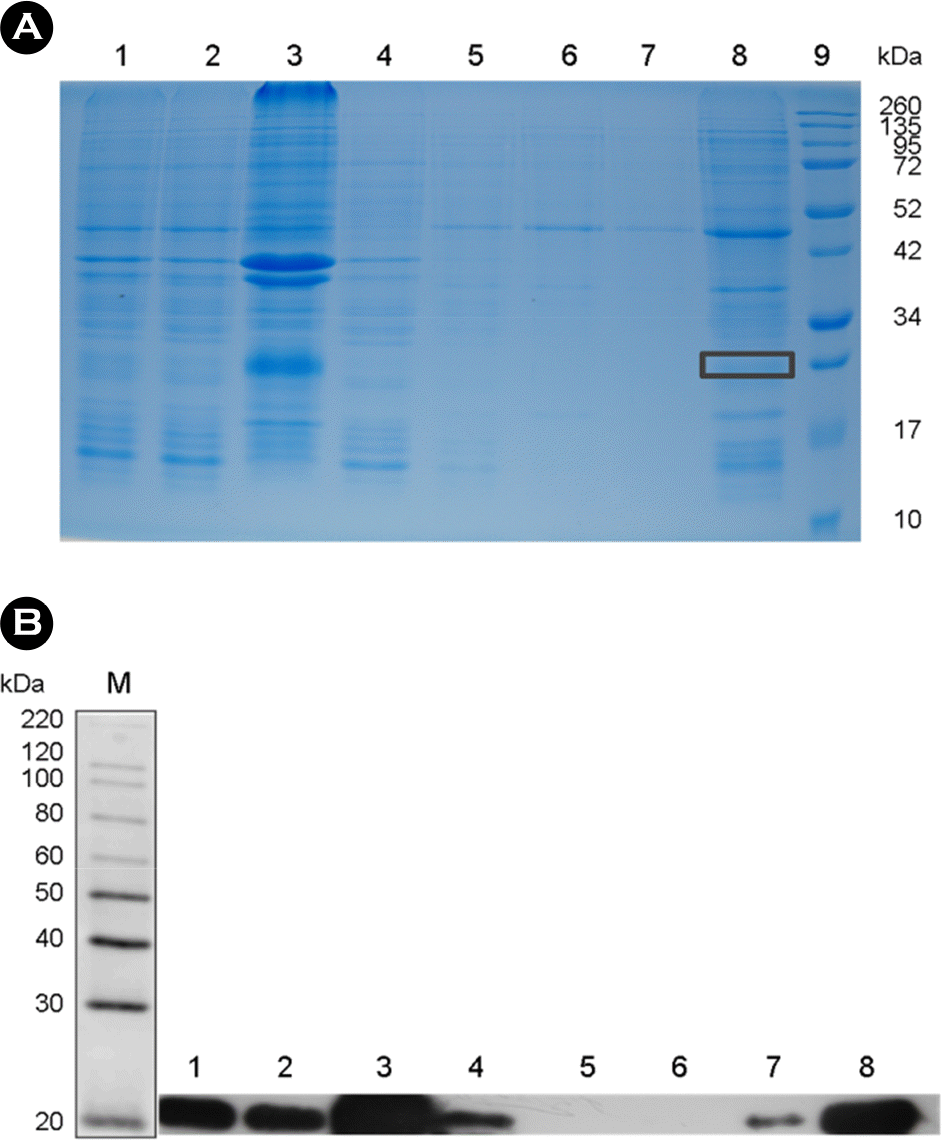 | Figure 2.Purification of recombinant NSP4 after the suspension and dissolution of inclusion bodies. (A) 12% SDS-PAGE of the fraction obtained by Ni-ion affinity chromatography; lane 1: total denatured lysates; lane 2, denatured lysis supernatant; lane 3, precipi-tate; lane 4, flow-through; lane 5~6, sequential column washings; lane 7, elution of the recombinant NSP4 protein from the column; lane 8, concentration of the recombinant NSP4 protein; and lane 9, protein molecular weight marker (Invitrogen, USA). (B) Western blotting using the monoclonal anti-His antibody of each fraction obtained during purification. Lane 1~8, as in SDS PAGE (panel A). |
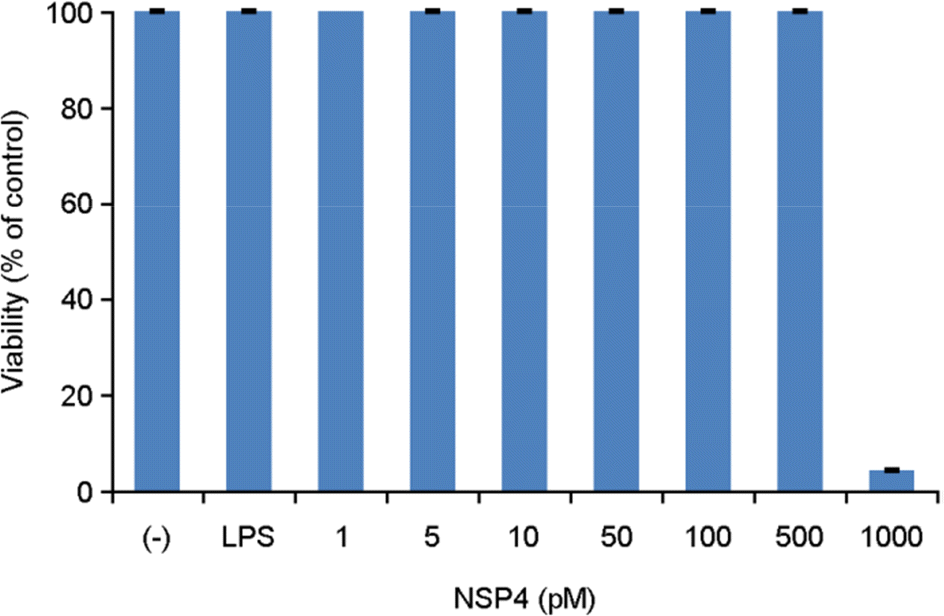 | Figure 3.Viability of RAW 264.7 cells after 24-h induction with NSP4. Viability was determined using the colorimetric MTT assay. |
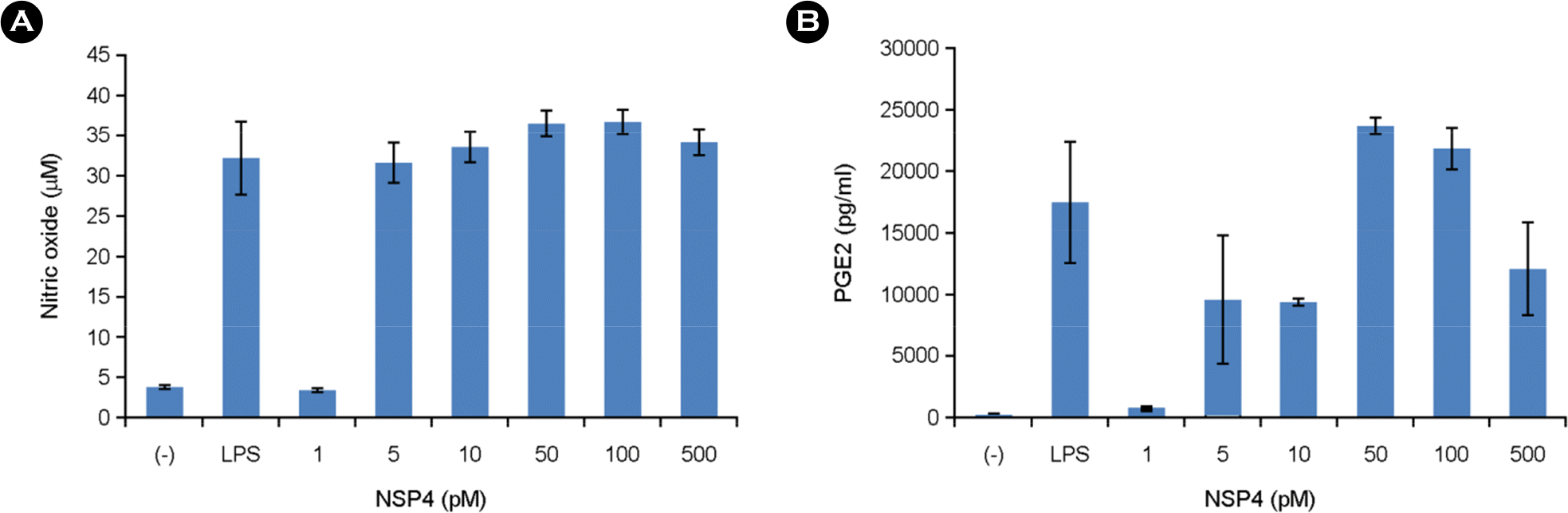 | Figure 4.Effects of NSP4 on the levels of NO and PGE2 in RAW 264.7 cells. Cells were treated for 24 h with 1, 5, 10, 50, 100, and 500 pM of various NSP4s. The concentrations of nitrite and PGE2 were measured as described in the Materials and Methods. The results are expressed as the mean ± standard error from three independent experiments. |
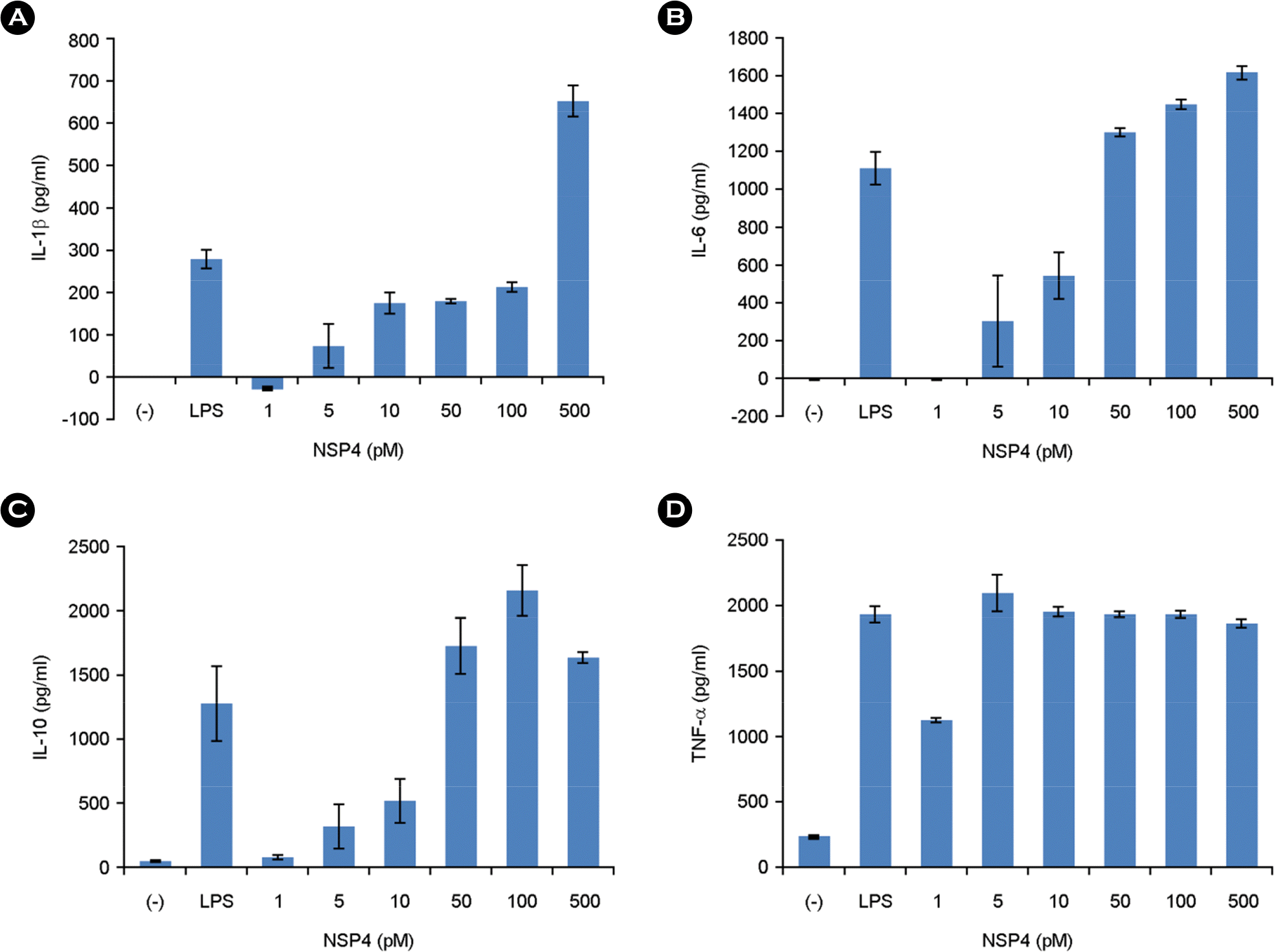 | Figure 5.Effect of NSP4 on the levels of IL-1β, IL-6, IL-10, and TNF-α in RAW 264.7 cells. The concentrations of IL-1β, IL-6, IL-10, and TNF-α released into the medium were determined by performing ELISA with the culture supernatant. They were measured as described in the Materials and Methods. Error bars shows the mean ± standard deviation of three measurements. |




 PDF
PDF ePub
ePub Citation
Citation Print
Print


 XML Download
XML Download