Abstract
Purpose
To analyze the correlation of polymorphisms of toll-like receptor 7 (TLR7) (rs179009) and toll-like receptor 9 (TLR9) (rs187084) in hepatitis C virus (HCV) infections in the Han population.
Materials and Methods
The genotypes of TLR7IVS2-151 in HCV infection were detected by Sanger sequencing using polymerase chain reaction-restriction fragment length polymorphism to determine the TLR9 T-1486C single nucleotide polymorphisms (SNP) for all enrolled patients.
Results
We found no significant difference between males with spontaneous clearance of HCV versus those chronically infected [χ2=2.71, p=0.10, odd ratios (OR)=0.58, 95% confidence interval (CI) 0.31-1.11]. However, significant differences were found for the distribution of TLR7 (rs179009) in females (χ2=9.46, p=0.01). In females, a significant difference was also found between chronic hepatitis C and those with spontaneous clearance of HCV in terms of TLR7 IVS2-151G/A allele frequencies (χ2=9.50, p=0.00, OR=0.46, 95% CI 0.28-0.75). In HCV-infected patients, no significant association was found between the frequency of TLR9 genotypes and alleles.
Conclusion
The site of TLR7 IVS2-151 (rs179009) G/A may be a factor for susceptibility of chronic HCV in the female Han population. TLR9T-1486C (rs18084) SNP may not play a major role in HCV infection. However, individual risk profiles for HCV infection did vary by sex and this relationship should be further investigated.
After discovery of the hepatitis C virus (HCV) more than 20 years ago, HCV infection has become a global problem which need to take a wide range of measures to control and prevention. There is a significant association between chronic hepatitis C (CHC) and the development of cirrhosis and hepatocellular carcinom in majority of the world. Approximately, 160 million chronically infected individuals and about 500000 related deaths occur in indusrtilized regions and an increasing threat in less-industrialized countries.2 Toll-like receptors (TLRs), belonging to a family of pathogen recognition receptors, are an essential part of the innate immune response and can detect conserved pathogen-associated molecular pattern (PAMPs) of bacteria, parasites, fungi, protozoa components, and viruses.3 The nucleotide-sensing TLRs include TLR3, TLR7, TLR8, and TLR9. TLR3 recognizes double-stranded RNA (dsRNA). TLR7 and TLR8 recognize single-stranded RNA (ssRNA), while TLR9 detects unmethylated CpG-containing DNA.4 TLRs are expressed by a variety of immune and non-immune cells, such as B lymphocytes, T lymphocytes, antigen-presenting cells, and fibroblastic synoviocytes.5 As well, TLR signals have been discovered in hepatitis B, hepatitis C, alcoholic liver diseases, non-alcoholic liver diseases, primary biliary cirrhosis, and more.6
TLR7 is mainly expressed in the endosome-lysosome membrane of plasmacytoid dendritic cells (pDCs), hepatic natural killer cells, and B lymphocytes. When the phagocytes take up a virus or virus-infected apoptotic cell, phogolysosome will degrade enzymes to relase viral RNA, Leading to ssRNA release and recognition by TLR7.7 TLR7 is interesting in regards to HCV-infection, because its engagement leads to production of increased levels of interferon-α.8,9 Zhang, et al.10 previously demonstrated that the HCV-specific G/U fragment is a motif sequence, recognized by TLR7 as a PAMP. The requirement for TLR7 to recognize HCV RNA was confirmed using specific inhibitors, RNAi, and by TLR7 overexpression. In addition, RNA instability, which reduces TLR7 expression, was also shown to be directly correlated with HCV replication and alterations in TLR7-induced interferon regulation factor (IRF)7-mediated cell activation. Furthermore, TLR7 has been shown to play a role in HCV-mediated evasion of host immune surveillance.11 Recently, stimulation of TLR7 with the investigational drug isatoribine was shown to lead to suppression of HCV-RNA in patients with chronic HCV-infection.12
TLR9 is considered as an immune mediator candidate in HBV because it is expressed in pDCs, binds cytidine-phosphate DNA motifs that are present in viruses, and stimulates the secretion of interferon-α when activated.13 The expression of TLR9 mRNA is elevated after stimulation of BV2 cells by HCV-positive serum. As well, TLR9 may increase the secretions of Th1 and Th2 cytokines so as to participate in the early inherent immune response during infection of the central nervous system by HCV.14 Data also suggest that TLR9 mRNA and protein are down-regulated in peripheral blood mononucleated cells of HCV-infected patients; they are also negatively correlated with serum viral copies and play an important role in detecting viral replication of HCV. Moreover, TLR9 stimulation shows antiviral effects in HCV-infected individuals;15 however, binding to TLR9 has only been demonstrated for DNA viruses and binding of HCV to TLR9 is unlikely.16
Large data supporting associations between single nucleotide polymorphisms (SNP) in TLRs and increased risk of bacteria, autoimmunity disease, and viral infection are being reported. Several studies have demonstrated an association between TLR7 gene polymorphisms and infection diseases in different populations; however, only one study showed that the TLR7 SNP rs179009 is related to susceptibility to HCV infection in Taiwanese patients in China.17 Most TLR SNPs, except TLR9 SNPs, have been investigated in terms of their relationship with chronic HCV.18 In the present study, we assessed the frequencies of TLR7 (rs179009) and TLR9 (rs187084) SNPs, in order to clarify the role of TLR7, TLR9 polymorphism in HCV infection.
One hundred and fifty (70 females and 80 males) chronic genotype 1 HCV-infected patients who visited Renmin Hospital of Wuhan University, Hubei Province, China, from January to June 2011, were enrolled in this study. Individuals with chronic infection exhibited anti-HCV antibody and HCV-RNA in serum >100 IU/mL before receiving any therapy. All patients had been infected with HCV for over 6 months, but were negative for HBsAg and anti-HIV. An additional 161 patients (75 females and 86 males) who exhibited spontaneous clearance of HCV patients showed anti-HCV antibody and undetectable HCV-RNA in serum (<100 IU/mL). All patients were native Chinese living in the central part of China. None of the subjects had a history of alcohol or drug abuse. Patients did not receive antiviral or immunosuppressive therapy prior to or during the course of the study. Subjects from this study were not statistically different in age or gender. This study was approved by the Ethics Committee of the Renmin Hospital of Wuhan University, and all subjects provided informed consent. The characteristics of the subjects are listed in Table 1.
We isolated genomic DNA from ethylene diamine tetraacetic acid (EDTA)-blood using Genomic DNA Extraction Kits (SBS Biological Engineering Co., Ltd, Shanghai, China) according to the manufacturer's instructions.
Polymorphisms of IVS2-151 (rs179009) in the TLR7 gene were detected by gene-sequencing. A 388 base-pair fragment containing the polymorphism site was amplified using TLR7-specific primers (forward 5'-GGAGTTTGGAAAT TAGGATTATGTT-3', reverse 5'-ACTTTGGCAGTG AATCTATGGC-3). Polymerase chain reaction (PCR) was performed using 2×mixture containing 1.5 mM MgCl2, 10 mM dNTPS, 2.5 U Taq DNA polymerase (Takara Biotechnology Co., Ltd, Dalian, China), 50 ng genomic DNA, and 20 µmol of each primer at a volume of 50 µL. The amplification conditions were as follows: denaturation at 94℃ for 5 min, followed by 35 cycles of denaturation at 94℃ for 45 s, annealing at 57℃ for 45 s, and extension at 72℃ for 1 min, with a final extension at 72℃ for 5 min. The PCR products were purified using Axygen quick PCR purification kit (Axyen Biotechnology Co., Ltd, Hangzhou, China) following the manufacturer's instructions. Subsequently, sequencing was performed using the Bigdye terminator version 3.1 cycle sequencing kit (ABI, Foster City, CA, USA). Following the manufacturer's instructions on an ABI 3100 xl Analyzer (ABI, Foster City, CA, USA), the results were analyzed with ABI PRISM® Seq software® version 2.0 (ABI, Foster City, CA, USA).
Polymorphisms of T-1486C (rs187084) in the TLR9 gene were detected by PCR restriction fragment length polymorphism (RFLP) analysis. A 419 base-pair fragment containing the polymorphic site was amplified using TLR9-specific primers (forward 5'-CATTCATTCAGCCTTCACTC-3'; reverse 5'-AT GTGCTGTTCCCTCTGC-3'). PCR was performed using a 2×mixture containing 1.5 mM MgCl2, 10 mM dNTP, 50 ng genomic DNA, 20 µmol of each primer, and 2.5 U Taq DNA polymerase (Takara Biotechnology Co., Ltd, Dalian, China) at a volume of 50 µL. Amplification was performed for 30 cycles with preheating at 95℃ for 5 min, followed by denaturation at 94℃ for 30 sec, annealing at 58.5℃ for 45 sec, and extension at 72℃ for 45 sec. The PCR product was incubated with the restriction enzyme AfI II (Lanheng Biotechnology Co., Ltd, Wuhan, China) for 1 h at 37℃, and the digestion products were resolved on a 3% agarose gel stained with ethidium bromide. The variant C allele produced a fragment of 419 bp; the wild-type T allele generated two fragments of 147 bp and 268 bp; and the variant T allele produced three fragments of 419 bp, 147 bp, and 268 bp. Additionally, all genotypes of TLR9 were further confirmed by direct sequencing of 20 randomly selected samples using the ABI 3100 xl Analyzer (ABI, Foster City, CA, USA).
The statistical package for Social Science version 13 (SPSS 13.0) for Windows (SPSS Inc., Chicago, IL, USA) was used for the statistical analyses. The genotype distribution of the SNPs of TLR7 and TLR9 were tested for departure from the Hardy-Weinberg equilibrium by means of χ2 analysis. Statistical significance in the differences in TLR genotypes and the alleles of included cases were calculated by Pearson χ2 test. Odd ratios (OR) and 95% confidence intervals (95% CIs) were calculated. All p-values <0.05 were considered statistically significant.
In total, 150 cases of chronic HCV and 161 cases of spontaneous clearance of HCV were enrolled. Clinical details and the results of biochemical analysis for the subjects at the time of the study are summarized in Table 1.
Distribution frequencies for different genotypes in the studied patients were compatible with the Hardy-Weinberg law. As well, Sanger sequencing was appropriate for determining the polymorphisms on TLR7 gene (Fig. 1). Table 2 shows the frequencies of genotypes of TLR7 (rs179009) in the chronic HCV infected and spontaneous clearance of HCV patients. Since TLR7 is located on an X chromosome, female and male individuals needed to be analyzed separately. In males, no significant association was found between the spontaneous clearance of HCV group and chronic HCV-infected group [χ2=2.71, p=0.10, OR=0.58, 95% CI (0.31-1.11)]. Among female patients in the spontaneous clearance of HCV group, 14 patients (18.67%) exhibited the GG genotype, 27 (36.00%) patients had the AA genotype, and 34 patients (45.33%) had the GC genotype. Among female patients in the chronic HCV-infected group, 7 patients (10.00%) demonstrated the GG genotype, 43 patients (61.43%) had the AA genotype, and 20 patients (28.57%) had the GA genotype. There were significant differences in the distribution of the TLR7 (rs179009) G/A polymorphism between the spontaneous clearance of HCV and the chronic HCV-infected groups in females (χ2=9.46, p=0.01). Pearson χ2 test showed that the AA genotype for TLR7 polymorphism in females could be a risk factor for chronicity of HCV infection [χ2=9.38, p=0.00, OR=0.35, 95% CI (0.18-0.69)], compared with GA and GG genotypes, whereas the AG genotype in females appears as a protective factor for chronicity of HCV infection [χ2=4.61, p=0.03, OR=2.12, 95% CI (1.06-4.21)], compared with AA and GG genotypes. A significant difference was also found in the comparison of the CHC and spontaneous clearance of HCV groups in terms of TLR7 IVS2-151G/A allele frequencies [χ2=9.50, p=0.00, OR=0.46, 95% CI (0.28-0.75)] in females.
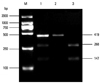
The PCR-RFLP method is proper for detecting polymorphisms on TLR9, which was genotyped in HCV infected patients (Fig. 2). The frequencies of genotypes of TLR9 (rs187084) in the chronic HCV and spontaneous clearance of HCV groups are shown in Table 3. No significant differences in the frequencies of genotypes were found between the chronic hepatitis C patients and spontaneous clearance of HCV patients. Furthermore, we analyzed the relationship between the two groups in allelic frequencies of TLR9 (rs187084), and there was no significant association in frequency between them [χ2=0.08, p=0.78, OR=1.05, 95% CI (0.76-1.43)]. Given that regulation of innate immune response is gender dependent,19 we analyzed the TLR9 polymorphism in males and females, but no statistical differences were found in the distribution frequencies of alleles or genotypes between males and females of the two groups.
No significant associations were observed between alanine aminotransferase (ALT), aspartate aminotransferase (AST), gamma-glutamyltransferase (GGT) and TLR7 or TLR9 genotypes in chronic HCV infection and spontaneous clearance of HCV patients (p>0.05).
A major function of TLRs is to recognize molecular patterns and evoke efficient and immediate immune responses appropriate to the nature of the bacterial, fungal, and viral infections.20 Human have 10 TLRs, distinct of them are activated in response to a given pathogen. For example, TLR2 forms a heterodimer with either TLR1 or TLR6 and binds bacterial lipoproteins. TLR4 recognizes the lipopolysaccharide of the outer membrane of Gram-negative bacteria.21 TLR3 recognizes dsRNA, polyinosinic-polycytidylic acid (polyI:C), genomic RNA purified from dsRNA viruses such as retrovirus, and dsRNA produced in the course of replication of ssRNA viruses.22 each TLR constitutes a class of membrane-bound pattern recognition receptors capable of detecting viral infections. TLR genetic variants and their downstream signaling molecules can influence the ability of affected individuals to respond adequately to TLR ligands, which can result in their altered susceptibility to HCV infection.23 We examined the effects of TLR7 and TLR9 polymorphisms in HCV infected patients. The major findings of this study were the TLR7 IVS2-151G/A polymorphism appears to be associated with gender in HCV infection. In females, this site mutation was associated with chronic infection susceptibility in patients who had been infected by HCV. We did not find any association between TLR9 polymorphism genotypes and HCV infection in the Chinese population.
The TLR7 gene is located on the X-chromosome, spanning three exons.24 The single nucleotide polymorphism at IVS2-151 of TLR7 was identified by Cheng, et al.25 The polymorphism, TLR7 IVS2-151G>A, changes the -151 nucleotide of the second intervening sequence from G to A and its function is still unclear. As HCV is a ssRNA virus, TLR7 have been thought to play a role in the immune response against HCV. In one study, TLR7 was revealed to be pivotal in HCV infection, as it can bind ssRNAs and lead to the production of large amounts of the antiviral cytokine interferon-α via dendritic cells.26 The pathway of TLR7 results in translocation of IRFs and nuclear NF-κB to the nucleus where IRF stimulates expression of type I IFN and a multitude of IFN-stimulated gene products, including enzymes, transcription factors, chemokines and cytokines. The signaling receptor TLR7 is directly critical for the efficient control of virus infections.27,28 HCV-infected cells can also directly activate pDCs to produce IFN-I via a TLR7-mediated pathway.29 Schott, et al.30 assessed the role of TLR7 polymorphism in 978 patients with chronic HCV infection and in 203 healthy controls, and they found that TLR7 polymorphisms at position 32 and 2403 were associated with chronic hepatitis C. TLR7 variants were also associated with response to IFN-α therapy. In addition, they also found that the c.1-120G TLR7 allele offers protection from the development of inflammation and fibrosis in male patients with chronic HCV-infection.31 Our results are different from that discovered by Wang, et al.17 The different results may be caused by the subjects from different areas, as well as differences in gender and disease groups. Many studies have shown that the regulation of innate immune responses is gender dependent: for example, men are more prone to infection, while women are more likely to suffer from autoimmune disease.32 In 2001, The Institute of Medicine suggested that assessments of differences in the incidence of many infectious diseases between males and females should also take into account differences in disease exposure.33 Thus, the patient's sex needs to be taken into account when future individual risk profiles for HCV infection are generated, which may give some clues in the study of the pathogenesis of persistent HCV infection.
The TLR9 gene is located on chromosome 3p21.3 and spans approximately 5 kb. TLR9 gene has two exons, and the second one is the major coding region.24 TLR9 -1237T/C (rs178084) is located within the putative promoter region that may influence transcriptional regulation of the TLR9 gene. Although TLR stimulation shows antiviral effects in HCV-infection, TLR9 has only been demonstrated to bind to DNA virus, and the binding of HCV to TLR9 is unlikely.31 Numerous studies have investigated the effect of TLR9 (rs187084) on diseases, such as rheumatoid arthritis,34 HBV,35 and so on. However, most studies failed to show any significant associations, except in cervical cancer; one previous study found that rs187084 (T-1486C) affected susceptibility to cervical cancer in Chinese women.36 In the present study, we set out to demonstrate associations between TLR9 polymorphism and the chronicity of HCV infection. However, no significant association in regards to genotype, allele frequency, and gender between patients with chronic HCV and spontaneous clearance of HCV subjects were found. Our results suggest that the TLR9 gene may not play a major role in persistent infection.
Some limitations in our study need to be addressed. First, all chronic HCV infection subjects were studied in-hospital, and thus may not be representative of the target population and inherent selection bias cannot be excluded. Second, the sample number in this study was moderate; other large population-based studies are needed to validate the findings. Third, only one SNP of TLR7/TLR9 was studied; therefore, we could not comprehensively investigate the association between SNPs of TLR7 and TLR9 and HCV infection susceptibility.
Figures and Tables
Fig. 1
Genotyping of the TLR7 gene polymorphism by singer sequencing. (A) The genotype is AA. (B) The genotype is GA. (C) The genotype is GA. TLR, toll-like receptor.

Fig. 2
Genotyping of the TLR9. M means DNA maker, lane 1 CT are digested in to three fragments, lane 2 CC are undigested, lane 3 digested into two fragments. TLR, toll-like receptor.
ACKNOWLEDGEMENTS
We thank Dr. Wei Wu for suggestions in reviewing the manuscript. This work was supported financially by the National Clinical Key Specialty Construction Projects.
References
2. Klevens RM, Hu DJ, Jiles R, Holmberg SD. Evolving epidemiology of hepatitis C virus in the United States. Clin Infect Dis. 2012; 55:Suppl 1. S3–S9.

4. Iwasaki A, Medzhitov R. Toll-like receptor control of the adaptive immune responses. Nat Immunol. 2004; 5:987–995.

5. Arpaia N, Barton GM. Toll-like receptors: key players in antiviral immunity. Curr Opin Virol. 2011; 1:447–454.

6. Testro AG, Visvanathan K. Toll-like receptors and their role in gastrointestinal disease. J Gastroenterol Hepatol. 2009; 24:943–954.

7. Askar E, Ramadori G, Mihm S. Toll-like receptor 7 rs179008/Gln11Leu gene variants in chronic hepatitis C virus infection. J Med Virol. 2010; 82:1859–1868.

8. Heil F, Hemmi H, Hochrein H, Ampenberger F, Kirschning C, Akira S, et al. Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. Science. 2004; 303:1526–1529.

9. Ito T, Amakawa R, Kaisho T, Hemmi H, Tajima K, Uehira K, et al. Interferon-alpha and interleukin-12 are induced differentially by Toll-like receptor 7 ligands in human blood dendritic cell subsets. J Exp Med. 2002; 195:1507–1512.

10. Zhang YL, Guo YJ, Bin Li, Sun SH. Hepatitis C virus single-stranded RNA induces innate immunity via Toll-like receptor 7. J Hepatol. 2009; 51:29–38.

11. Chang S, Kodys K, Szabo G. Impaired expression and function of toll-like receptor 7 in hepatitis C virus infection in human hepatoma cells. Hepatology. 2010; 51:35–42.

12. Horsmans Y, Berg T, Desager JP, Mueller T, Schott E, Fletcher SP, et al. Isatoribine, an agonist of TLR7, reduces plasma virus concentration in chronic hepatitis C infection. Hepatology. 2005; 42:724–731.

13. Szabo G, Dolganiuc A, Mandrekar P. Pattern recognition receptors: a contemporary view on liver diseases. Hepatology. 2006; 44:287–298.

14. Hu K, Wang GW, Wang ZH. [TLR9 mRNA expression and tumor necrosis factor-alpha/interleukin-6 secretion in murine microglia by hepatitis C virus stimulation]. Zhonghua Yi Xue Za Zhi. 2011; 91:1070–1074.
15. Broering R, Wu J, Meng Z, Hilgard P, Lu M, Trippler M, et al. Toll-like receptor-stimulated non-parenchymal liver cells can regulate hepatitis C virus replication. J Hepatol. 2008; 48:914–922.

16. Akira S, Uematsu S, Takeuchi O. Pathogen recognition and innate immunity. Cell. 2006; 124:783–801.

17. Wang CH, Eng HL, Lin KH, Chang CH, Hsieh CA, Lin YL, et al. TLR7 and TLR8 gene variations and susceptibility to hepatitis C virus infection. PLoS One. 2011; 6:e26235.

18. Sawhney R, Visvanathan K. Polymorphisms of toll-like receptors and their pathways in viral hepatitis. Antivir Ther. 2011; 16:443–458.

19. Fish EN. The X-files in immunity: sex-based differences predispose immune responses. Nat Rev Immunol. 2008; 8:737–744.

20. Song DH, Lee JO. Sensing of microbial molecular patterns by Toll-like receptors. Immunol Rev. 2012; 250:216–229.

21. Kang JY, Lee JO. Structural biology of the Toll-like receptor family. Annu Rev Biochem. 2011; 80:917–941.

22. Sato A, Iwasaki A. Induction of antiviral immunity requires Toll-like receptor signaling in both stromal and dendritic cell compartments. Proc Natl Acad Sci U S A. 2004; 101:16274–16279.

23. Wieland SF, Chisari FV. Stealth and cunning: hepatitis B and hepatitis C viruses. J Virol. 2005; 79:9369–9380.

24. Du X, Poltorak A, Wei Y, Beutler B. Three novel mammalian toll-like receptors: gene structure, expression, and evolution. Eur Cytokine Netw. 2000; 11:362–371.
25. Cheng PL, Eng HL, Chou MH, You HL, Lin TM. Genetic polymorphisms of viral infection-associated Toll-like receptors in Chinese population. Transl Res. 2007; 150:311–318.

26. Colonna M, Krug A, Cella M. Interferon-producing cells: on the front line in immune responses against pathogens. Curr Opin Immunol. 2002; 14:373–379.

27. García-Sastre A, Biron CA. Type 1 interferons and the virus-host relationship: a lesson in détente. Science. 2006; 312:879–882.
28. Gale M Jr, Foy EM. Evasion of intracellular host defence by hepatitis C virus. Nature. 2005; 436:939–945.

29. Takahashi K, Asabe S, Wieland S, Garaigorta U, Gastaminza P, Isogawa M, et al. Plasmacytoid dendritic cells sense hepatitis C virus-infected cells, produce interferon, and inhibit infection. Proc Natl Acad Sci U S A. 2010; 107:7431–7436.

30. Schott E, Witt H, Neumann K, Bergk A, Halangk J, Weich V, et al. Association of TLR7 single nucleotide polymorphisms with chronic HCV-infection and response to interferon-a-based therapy. J Viral Hepat. 2008; 15:71–78.

31. Schott E, Witt H, Neumann K, Taube S, Oh DY, Schreier E, et al. A Toll-like receptor 7 single nucleotide polymorphism protects from advanced inflammation and fibrosis in male patients with chronic HCV-infection. J Hepatol. 2007; 47:203–211.

33. Exploring the biological contributions to human health: does sex matter? J Womens Health Gend Based Med. 2001; 10:433–439.
34. Jaen O, Petit-Teixeira E, Kirsten H, Ahnert P, Semerano L, Pierlot C, et al. No evidence of major effects in several Toll-like receptor gene polymorphisms in rheumatoid arthritis. Arthritis Res Ther. 2009; 11:R5.





 PDF
PDF ePub
ePub Citation
Citation Print
Print




 XML Download
XML Download