Abstract
Background
We investigated the prevalence of plasmid-mediated quinolone resistance and its association with extended-spectrum beta-lactamase (ESBL) and AmpC beta-lactamase in Enterobacteriaceae.
Methods
A total of 347 non-duplicated isolates of Enterobacteriaceae were collected between August and October 2006 from 2 hospitals. Qnr determinant screening was conducted using PCR amplification, and all positive results were confirmed by direct sequencing. Qnr-positive strains were determined on the basis of the presence of ESBL and AmpC beta-lactamase genes.
Results
The qnr gene was detected in 47 of 347 clinical Enterobacteriaceae isolates. Among the 47 qnr-positive strains, Klebsiella pneumoniae (N=29) was the most common, followed by Escherichia coli (N=6), Enterobacter cloacae (N=6), Citrobacter freundii (N=5), and Enterobacter aerogenes (N=1). These isolates were identified as qnrA1 (N=6), 8 qnrB subtypes (N=40), and qnrS1 (N=1). At least 1 ESBL was detected in 38 of the 47 qnr-positive strains. Qnr-positive strains also showed high positive rates of ESBL or AmpC beta-lactamase, such as TEM, SHV, CTX-M, and DHA. DHA-1 was detected in 23 of 47 qnr-positive strains, and this was co-produced with 1 qnrA1 and 22 qnrB4. Strains harboring MIR-1T and CMY were also detected among the qnr-positive strains. Antimicrobial-resistance rates of qnr-positive strains to ciprofloxacin, levofloxacin, norfloxacin, nalidixic acid, and moxifloxacin were 51.1%, 46.8%, 46.8%, 74.5%, and 53.2%, respectively.
The first plasmid-mediated quinolone resistance determinant, qnrA, was reported in 1998 in a Klebsiella pneumoniae strain from the United States [1]. Since then, 2 additional qnr determinants, qnrB and qnrS, have been identified in Enterobacteriaceae species [2, 3]. These determinants increase the minimum inhibitory concentrations (MICs) of quinolones by 8- to 32-fold, preventing the inhibition of DNA gyrase [4]. These 3 major groups of qnr determinants have since been detected in various Enterobacteriaceae species worldwide [5-9], and many qnr variants are listed in GenBank. Among them, qnrB sequences are particularly numerous; hence, qnrB sequences with the same allele number and dissimilar sequences submitted by different investigators or with the same sequences and different allele numbers can be found [10]. This situation results in confusion for many investigators when numbering their qnr sequences, particularly qnrB variants; thus, Jacoby et al. [10] adjusted all previously submitted sequences and proposed a new, well-organized numbering system for qnr alleles.
Association of qnr genes with extended-spectrum beta-lactamases (ESBLs) or AmpC beta-lactamases is also noteworthy; some strains containing a qnrA gene express class A or class C beta-lactamase, such as SHV, CTX, VEB-1, or DHA-1 [11, 12]. Some reports show an association of qnrB with an ESBL and AmpC beta-lactamase [3, 13].
Although there are some reports examining the prevalence of qnr determinants in Korea, these were restricted to specific bacterial species such as Escherichia coli, K. pneumoniae, Enterobacter cloacae, Citrobacter freundii, and Serratia marcescens regardless of the type of specimen [8, 13-16]. Additionally, there are few data regarding the differences in qnr prevalence according to the type of clinical specimen and bacterial species and its association with various ESBLs and AmpC beta-lactamases. The aims of this study were to investigate the prevalence of 3 groups of plasmid-mediated quinolone resistance determinants (qnrA, qnrB, and qnrS) and to examine the association of these determinants with ESBLs and AmpC beta-lactamases in Enterobacteriaceae.
A total of 347 non-duplicated clinical strains of Enterobacteriaceae were collected from various clinical specimens from 2 hospitals, namely, the 902-bed tertiary-care Busan Paik Hospital (N=267) and the 200-bed secondary-care Dongrae Paik Hospital (N=80) in 2006. Sampling locations included urine (N=150), sputum or bronchial washings (N=59), abscess or wound pus (N=52), drainage discharge (N=30), peritoneal fluid (N=14), cervical discharge (N=10), gastric juice (N=6), blood culture (N=6), tissue (N=4), stool (N=4), bile (N=3), vaginal discharge (N=3), pleural fluid (N=2), catheter tip (N=2), intrauterine device (N=1), and cerebrospinal fluid (N=1). The strains were identified using routine laboratory protocols, including conventional biochemical tests and the VITEK system (bioMèrieux Vitek Inc., Hazelwood, MO, USA). These strains were identified as E. coli (N=159), K. pneumoniae (N=100), E. cloacae (N=30), S. marcescens (N=13), Proteus mirabilis (N=10), Morganella morganii (N=6), C. freundii (N=7), Enterobacter aerogenes (N=4), Klebsiella oxytoca (N=3), Citrobacter braakii (N=2), Proteus vulgaris (N=2), Proteus penneri (N= 2), Citrobacter amalonaticus (N=1), Enterobacter asburiae (N=1), Enterobacter hormaechei (N=1), Enterobacter intermedius (N=1), Escherichia fergusonii (N=1), Klebsiella rhinoscleromatis (N=1), Klebsiella species (N=1), Providencia rettgeri (N=1), and Providencia stuartii (N=1). All isolates were stored in skim milk at -70℃ until further molecular and microbiological evaluation.
Genomic DNA of the bacterial strains was amplified by PCR for the detection of 3 groups of qnr determinants (qnrA, qnrB, and qnrS) as described by Gay et al. [5]. All procedures for PCR and sequencing were the same as those described in our previous report [8]. Briefly, colonies were suspended in 50 µL of water in a microcentrifuge tube and boiled to prepare DNA templates. The primers used to amplify qnrA to yield a 516-bp product were 5'-ATTTCTCACGCCAGGATTTG-3' and 5'-GATCGGCAAAGGTTAGGTCA-3'. Those used for qnrB to yield a 469-bp product were 5'-GATCGTGAAAGCCAGAAAGG-3' and 5'-ACGATGCCTGGTAGTTGTCC-3'. Those used for qnrS to yield a 417-bp product were 5'-ACGACATTCGTCAACTGCAA-3' and 5'-TAAATTGGCACCCTGTAGGC-3'. The 3 primer pairs were added to a template and PCR premix (Bioneer, Daejeon, Korea). PCR conditions were 94℃ for 45 sec, 53℃ for 45 sec, and 72℃ for 60 sec, with the cycle repeated 32 times. Reaction mixtures without a DNA template served as negative controls. All positive amplicons were sequenced to confirm the subtypes of the 3 qnr genes. Amplified DNA was isolated using the Qiagen gel extraction kit (Qiagen Inc., Hilden, Germany), and sequencing was performed using the Big Dye Terminator Cycle Sequencing kit (Applied Biosystems, Foster City, CA, USA) and an ABI PRISM 3130xl genetic analyzer (Applied Biosystems). Both amplicon strands were sequenced using the same primers as for PCR amplification. We numbered the qnr allele subtypes according to the qnr gene nomenclature of Jacoby et al. [10].
Plasmid DNA was extracted using a Qiagen Plasmid Purification kit (Qiagen Inc.) following the manufacturer's recommendations to detect ESBL and AmpC beta-lactamase in qnr-positive strains. In previous experiments, this plasmid DNA was used as a template for PCR with a series of primers designed to detect class A beta-lactamase genes and their extended-spectrum derivatives (Table 1): blaCTX-M-1, blaCTX-M-2, blaCTX-M-8, blaCTX-M-9, blaPER-1, blaVEB, blaIBC/GES, blaTEM, and blaSHV [17, 18]. PCR products were subjected to direct sequencing, with both strands being sequenced to reduce error. We also investigated the presence of plasmid-mediated AmpC beta-lactamase among qnr-positive strains using the primers developed by Perez-Perez and Hanson for 6 groups: ACC, FOX, MOX/CMY, DHA, CIT, and EBC [19]. These groups could be used to detect the following target genes: blaDHA, blaMOX, blaCMY, blaLAT, blaBIL, blaACC, blaMIR, blaACT, and blaFOX. We used 2 multiplex PCR methods described by Lee et al. [20] that were modified from the method of Perez-Perez and Hanson [19] to amplify the plasmid-mediated AmpC beta-lactamase genes. We used additional primers for sequencing the plasmid-mediated AmpC beta-lactamase genes and identified them by sequencing the PCR-amplified products. Primers used for the detection and sequencing of ESBL and the plasmid-mediated AmpC beta-lactamase are shown in Table 1.
Susceptibility testing against quinolones was determined using the broth microdilution method recommended by the CLSI [21]. Isolates were tested against the following 5 agents in cation-adjusted Mueller-Hinton broth (BD, Sparks, MD, USA): ciprofloxacin (Sigma, Steinheim, Germany), levofloxacin (Sigma), norfloxacin (Sigma), nalidixic acid (Sigma), and moxifloxacin (Bayer AG, Wuppertal, Germany). MICs were determined after incubation at 35℃ for 24 hr in ambient air. E. coli ATCC 25922 was used as the control strain. The interpretive criteria were those published in the relevant CLSI document [22]. For other antimicrobial agents, susceptibility testing was in accordance with routine laboratory protocols performed using the VITEK system or the disk diffusion assay.
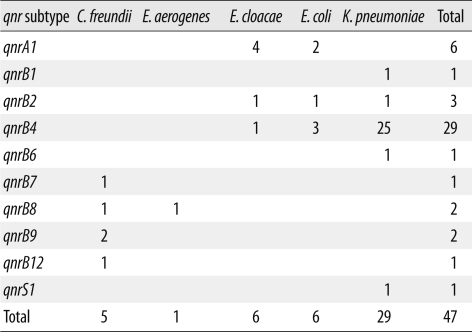
The qnr gene was detected in 47 of the 347 Enterobacteriaceae isolates by using PCR and sequencing. The positive rates were higher among organisms from the tertiary-care hospital (15.7%, N=42) than in the secondary-care facility (6.3%, N=5). The specimens showing qnr-positive results included sputum (N=15), urine (N=13), wound pus (N=10), drainage discharge (N=4), peritoneal fluid (N=2), cervical discharge (N=1), pleural fluid (N=1), and stool (N=1). Qnr genes were detected at a higher rate in respiratory specimens (15/59, 25.4%) than in urine (13/150, 8.7%). The most commonly observed qnr-positive pathogen was K. pneumoniae (N=29), followed by E. coli (N=6), E. cloacae (N=6), C. freundii (N=5), and E. aerogenes (N=1).
Ten qnr subtypes, including 1 qnrA, 8 qnrB, and 1 qnrS, were detected. There was a considerable difference in subtypes according to the specific species (Table 2). Thus, qnrA was primarily detected in E. cloacae and E. coli; on the other hand, qnrB was highly prevalent in K. pneumoniae. The qnrA gene was detected in 6 of the 47 qnr-positive isolates, and the sequence was identical for all strains. The qnrB gene was detected in 40 of the 47 qnr-positive isolates. These isolates showed considerable multiplicity of subtypes and sequences. The high rates of the qnrB4 subtype among qnrB subtypes (29/40) should be noted. In addition to these qnrB4 subtypes, 1 isolate containing qnrB1, 3 having qnrB2, 1 having qnrB6, 1 having qnrB7, 2 having qnrB8, 2 having qnrB9, and 1 having qnrB12 were detected. The qnrS1 gene was found in only 1 strain and exhibited 100% identity with the reference qnrS1 gene sequence.
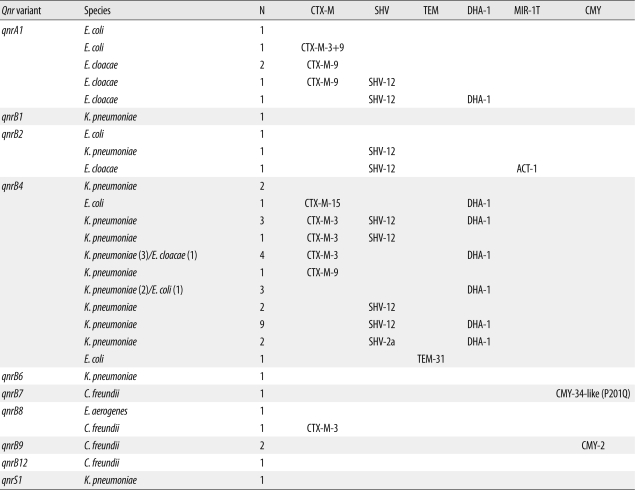
At least 1 ESBL or AmpC beta-lactamase was detected in 38 of the 47 qnr-positive isolates (Table 3). The beta-lactamase production rates of qnr-positive K. pneumoniae, E. coli, E. cloacae, and C. freundii were 82.8% (24/29), 66.7% (4/6), 100% (6/6), and 80.0% (4/5), respectively. No ESBLs were detected in E. aerogenes harboring qnrB8. The qnr-positive strains showed high positive rates of beta-lactamases such as TEM, SHV, CTX-M, and DHA. CTX-M-3, CTX-M-9, and SHV-12 were highly prevalent among ambler class A beta-lactamases. In plasmid-mediated AmpC beta-lactamase, DHA-1 was detected in 23 of 47 qnr-positive strains and was co-produced with 1 qnrA1 and 22 qnrB4. Strains harboring MIR-1T and CMY primers were also detected among qnr-positive isolates. There were no qnr-positive clinical isolates harboring MOX, ACC, FOX, PER-1, VEB, or GES.
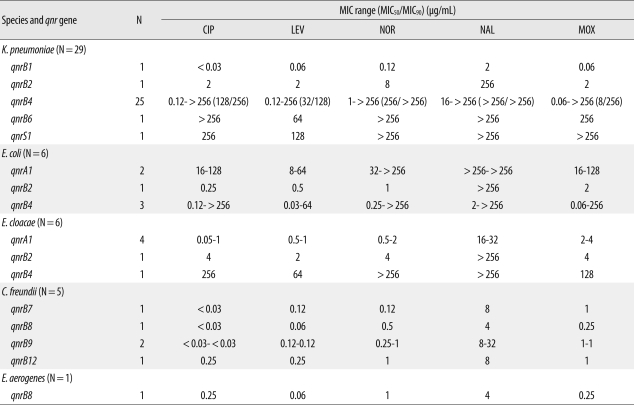
Of the 47 qnr-positive strains examined, the resistance rates to ampicillin, ampicillin-sulbactam, aztreonam, cefazolin, tobramycin, amikacin, ceftriaxone, cefepime, and trimethoprim-sulfamethoxazole were 100%, 84.8%, 69.6%, 87.0%, 69.6%, 43.5%, 45.7%, 34.8%, and 47.8%, respectively. The antimicrobial-resistance rates of qnr-positive strains to ciprofloxacin, levofloxacin, norfloxacin, nalidixic acid, and moxifloxacin were 51.1%, 46.8%, 46.8%, 74.5%, and 53.2%, respectively (Table 4). The plasmid-mediated quinolone resistance rate was very high in both quinolone-resistant (24/96) and quinolone-susceptible strains (22/248). No imipenem- or meropenem-resistant strains of Enterobacteriaceae were observed.
In this study, 47 of 347 (12.4%) Enterobacteriaceae showed qnr positive results. There was a considerable difference in qnr prevalence between the tertiary-care (15.7%) and secondary-care hospitals (6.3%) evaluated in this study. Generally, antimicrobial resistance rates are higher in tertiary-care hospitals [23, 24], and these differences are thought to be applicable to qnr-positive rates. Although several articles regarding qnr genes have been published, they focus on a specific species and do not report specimen types. In the present study, we isolated qnr-positive strains from various clinical specimens. We also observed a characteristically high qnr-positive rate of respiratory specimens (25.4%) compared with other specimens, including urine (8.7%), wound pus (19.2%), discharge (13.3%), and peritoneal fluid (14.3%). This may be because K. pneumoniae is highly prevalent in respiratory specimens. Because qnr-positive strains can spread in hospital environments and cause fatal nosocomial infection, the prevalence of qnr genes should be periodically investigated.
The numbers of qnrB subtypes were diverse compared with those found in our previous study [8]. Additionally, 6 qnrA strains were different from those in our previous study. We confirmed that qnrA1 (6/47) and qnrB4 (29/47) strains were common, with these 2 subtypes accounting for 74.5% (35/47) of the total qnr-positive strains.
The prevalence of the qnr gene was caused by qnrB4-positive K. pneumoniae in our hospital; a significant number of the qnr-positive strains were of this unique subtype. Although the rate of qnr in C. freundii (42.9%) was relatively high, the numbers of C. freundii were small. Few reports have investigated qnr prevalence in C. freundii; hence, further evaluation is necessary [16, 25].
The presence of qnr was significantly related to the co-production of ESBL or AmpC beta-lactamase. Surprisingly, at least 1 ESBL was detected in 38 of 47 qnr-positive isolates examined in this study. The 14, 21, and 3 qnr-positive strains showed co-production of ESBL or AmpC beta-lactamase in 1, 2, and 3 genes, respectively. This is primarily attributable to the qnrA1 and qnrB4 subtypes.
There are some reports of an association between qnr and ESBL. Wang et al. [11] reported that several qnrA-positive strains produced SHV-7 and CTX-M-9. In our study, CTX-M-9 was common (4 of 6) among qnrA1-positive strains, but 1 strain produced SHV-12 instead of SHV-7. Another report revealed positive results for both qnrA and VEB-1 in E. coli [6]. The investigators subsequently detected the qnrA gene in 11 of 23 blaVEB-1-positive Enterobacter sakazakii strains from Bangkok [26]. They also stated that the qnrA gene was not identified in positive enterobacterial isolates from any country except in previous 2 reports. No VEB-1-positive strain was identified among the 47 qnr-positive Enterobacteriaceae examined in our study.
In Korea, Pai et al. [13] described a clear association between qnrB4 and DHA-1 strains. In our study, many qnrB4-positive strains contained DHA-1 and, except for qnrB4 strains, only 1 strain produced DHA-1 in qnrA1 among all qnr subtypes. Thus, we presumed that an association exists between qnrB4 and DHA-1. However, this association does not appear to be limited to DHA-1. Although Pai et al. [13] stated that qnrB-positive strains were not closely related to SHV-12, we obtained different results. The co-production rates of SHV were very high in the qnrB4 strain. SHV was produced in 17 strains among the 29-qnrB4 positive strains. Additionally, 12 SHV were co-produced among 22 strains that were positive for both DHA-1 and qnrB4, and 20 of 23 DHA-1-positive strains harbored either SHV or CTX-M ESBLs. SHV-12 and DHA-1 were the most commonly observed strains in this study. In a previous report, CTX-M was found to be rare among qnr-positive strains, and these strains were primarily qnrA-positive ones [13]. However, 14 of 47 qnr-positive strains were CTX-M positive in the present study. These strains and 10 CTX-M strains, including 3 CTX-M-3, 1 CTX-M-9, and 1 CTX-M-15, harbored qnrB4. This was a characteristic finding in our series. Investigating the genetic environments, including CR1, to understand the mechanisms of association between qnr and ESBL is important. Additional evaluation is required to obtain a precise understanding of these mechanisms.
In the present study, we verified that the qnr genes were prevalent among quinolone-susceptible strains as well as among quinolone-resistant strains. This suggests that the qnr gene is widely dispersed among Enterobacteriaceae. This is because of the high prevalence of qnrB4 strains, which are closely related to various ESBL and plasmid-mediated AmpC beta-lactamases.
Acknowledgement
This work was supported by Basic Science Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (MEST) (KRF-2007-331-E00208).
References
1. Martinez-Martinez L, Pascual A, Jacoby GA. Quinolone resistance from a transferable plasmid. Lancet. 1998; 351:797–799. PMID: 9519952.
2. Hata M, Suzuki M, Matsumoto M, Takahashi M, Sato K, Ibe S, et al. Cloning of a novel gene for quinolone resistance from a transferable plasmid in Shigella flexneri 2b. Antimicrob Agents Chemother. 2005; 49:801–803. PMID: 15673773.
3. Jacoby GA, Walsh KE, Mills DM, Walker VJ, Oh H, Robicsek A, et al. qnrB, another plasmid-mediated gene for quinolone resistance. Antimicrob Agents Chemother. 2006; 50:1178–1182. PMID: 16569827.
4. Tran JH, Jacoby GA. Mechanism of plasmid-mediated quinolone resistance. Proc Natl Acad Sci USA. 2002; 99:5638–5642. PMID: 11943863.

5. Gay K, Robicsek A, Strahilevitz J, Park CH, Jacoby G, Barrett TJ, et al. Plasmid-mediated quinolone resistance in non-Typhi serotypes of Salmonella enterica. Clin Infect Dis. 2006; 43:297–304. PMID: 16804843.
6. Mammeri H, Van De Loo M, Poirel L, Martinez-Martinez L, Nordmann P. Emergence of plasmid-mediated quinolone resistance in Escherichia coli in Europe. Antimicrob Agents Chemother. 2005; 49:71–76. PMID: 15616277.
7. Poirel L, Leviandier C, Nordmann P. Prevalence and genetic analysis of plasmid-mediated quinolone resistance determinants QnrA and QnrS in Enterobacteriaceae isolates from a French university hospital. Antimicrob Agents Chemother. 2006; 50:3992–3997. PMID: 16982792.
8. Shin JH, Jung HJ, Lee JY, Kim HR, Lee JN, Chang CL. High rates of plasmid-mediated quinolone resistance QnrB variants among ciprofloxacin-resistant Escherichia coli and Klebsiella pneumoniae from urinary tract infections in Korea. Microb Drug Resist. 2008; 14:221–226. PMID: 18707554.
9. Wu JJ, Ko WC, Tsai SH, Yan JJ. Prevalence of plasmid-mediated quinolone resistance determinants QnrA, QnrB, and QnrS among clinical isolates of Enterobacter cloacae in a Taiwanese hospital. Antimicrob Agents Chemother. 2007; 51:1223–1227. PMID: 17242140.
10. Jacoby G, Cattoir V, Hooper D, Martinez-Martinez L, Nordmann P, Pascual A, et al. qnr gene nomenclature. Antimicrob Agents Chemother. 2008; 52:2297–2299. PMID: 18426894.
11. Wang M, Tran JH, Jacoby GA, Zhang Y, Wang F, Hooper DC. Plasmid-mediated quinolone resistance in clinical isolates of Escherichia coli from Shanghai, China. Antimicrob Agents Chemother. 2003; 47:2242–2248. PMID: 12821475.
12. Nordmann P, Poirel L. Emergence of plasmid-mediated resistance to quinolones in Enterobacteriaceae. J Antimicrob Chemother. 2005; 56:463–469. PMID: 16020539.
13. Pai H, Seo MR, Choi TY. Association of QnrB determinants and production of extended-spectrum beta-lactamases or plasmid-mediated AmpC beta-lactamases in clinical isolates of Klebsiella pneumoniae. Antimicrob Agents Chemother. 2007; 51:366–368. PMID: 17074790.
14. Jeong JY, Yoon HJ, Kim ES, Lee Y, Choi SH, Kim NJ, et al. Detection of qnr in clinical isolates of Escherichia coli from Korea. Antimicrob Agents Chemother. 2005; 49:2522–2524. PMID: 15917562.
15. Kim MH, Sung JY, Park JW, Kwon GC, Koo SH. Coproduction of qnrB and armA from extended-spectrum beta-lactamase-producing Klebsiella pneumoniae. Korean J Lab Med. 2007; 27:428–436. PMID: 18160833.
16. Park YJ, Yu JK, Lee S, Oh EJ, Woo GJ. Prevalence and diversity of qnr alleles in AmpC-producing Enterobacter cloacae, Enterobacter aerogenes, Citrobacter freundii and Serratia marcescens: a multicentre study from Korea. J Antimicrob Chemother. 2007; 60:868–871. PMID: 17660263.
17. Bae IK, Lee BH, Hwang HY, Jeong SH, Hong SG, Chang CL, et al. A novel ceftazidime-hydrolysing extended-spectrum beta-lactamase, CTX-M-54, with a single amino acid substitution at position 167 in the omega loop. J Antimicrob Chemother. 2006; 58:315–319. PMID: 16785225.
18. Ryoo NH, Kim EC, Hong SG, Park YJ, Lee K, Bae IK, et al. Dissemination of SHV-12 and CTX-M-type extended-spectrum beta-lactamases among clinical isolates of Escherichia coli and Klebsiella pneumoniae and emergence of GES-3 in Korea. J Antimicrob Chemother. 2005; 56:698–702. PMID: 16141280.
19. Pérez-Pérez FJ. Detection of plasmid-mediated AmpC beta-lactamase genes in clinical isolates by using multiplex PCR. J Clin Microbiol. 2002; 40:2153–2162. PMID: 12037080.
20. Lee K, Hong SG, Park YJ, Lee HS, Song W, Jeong J, et al. Evaluation of phenotypic screening methods for detecting plasmid-mediated AmpC beta-lactamases-producing isolates of Escherichia coli and Klebsiella pneumoniae. Diagn Microbiol Infect Dis. 2005; 53:319–323. PMID: 16269222.
21. Clinical and Laboratory Standards Institute. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically; approved standard-seventh edition, M7-A7. 2006. Wayne, PA: Clinical and Laboratory Standards Institute.
22. Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; Sixteenth informational supplement, M100-S16. 2006. Wayne, PA: Clinical and Laboraotory Standards Institute.
23. Yoo J, Sohn ES, Chung GT, Lee EH, Lee KR, Park YK, et al. Five-year report of national surveillance of antimicrobial resistance in Pseudomonas aeruginosa isolated from non-tertiary care hospitals in Korea (2002-2006). Diagn Microbiol Infect Dis. 2008; 60:291–294. PMID: 18068936.
24. Hong SG, Yong D, Lee K, Kim EC, Lee WK, Jeong SH, et al. Antimicrobial resistance of clinically important bacteria isolated from hospital located in representative provinces of Korea. Korean J Clin Microbiol. 2003; 6:29–36.
25. Jonas D, Biehler K, Hartung D, Spitzmüller B, Daschner FD. Plasmid-mediated quinolone resistance in isolates obtained in german intensive care units. Antimicrob Agents Chemother. 2005; 49:773–775. PMID: 15673764.

26. Poirel L, Van De Loo M, Mammeri H, Nordmann P. Association of plasmid-mediated quinolone resistance with extended-spectrum beta-lactamase VEB-1. Antimicrob Agents Chemother. 2005; 49:3091–3094. PMID: 15980408.
27. Song W, Kim JS, Kim HS, Yong D, Jeong SH, Park MJ, et al. Increasing trend in the prevalence of plasmid-mediated AmpC beta-lactamases in Enterobacteriaceae lacking chromosomal ampC gene at a Korean university hospital from 2002 to 2004. Diagn Microbiol Infect Dis. 2006; 55:219–224. PMID: 16545935.
28. Coudron PE, Hanson ND, Climo MW. Occurrence of extended-spectrum and AmpC beta-lactamases in bloodstream isolates of Klebsiella pneumoniae: isolates harbor plasmid-mediated FOX-5 and ACT-1 AmpC beta-lactamases. J Clin Microbiol. 2003; 41:772–777. PMID: 12574281.
29. Nadjar D, Rouveau M, Verdet C, Donay L, Herrmann J, Lagrange PH, et al. Outbreak of Klebsiella pneumoniae producing transferable AmpC-type beta-lactamase (ACC-1) originating from Hafnia alvei. FEMS Microbiol Lett. 2000; 187:35–40. PMID: 10828397.
Table 1
Primers used for the detection and sequencing of class A beta-lactamases and plasmid-mediated AmpC beta-lactamases
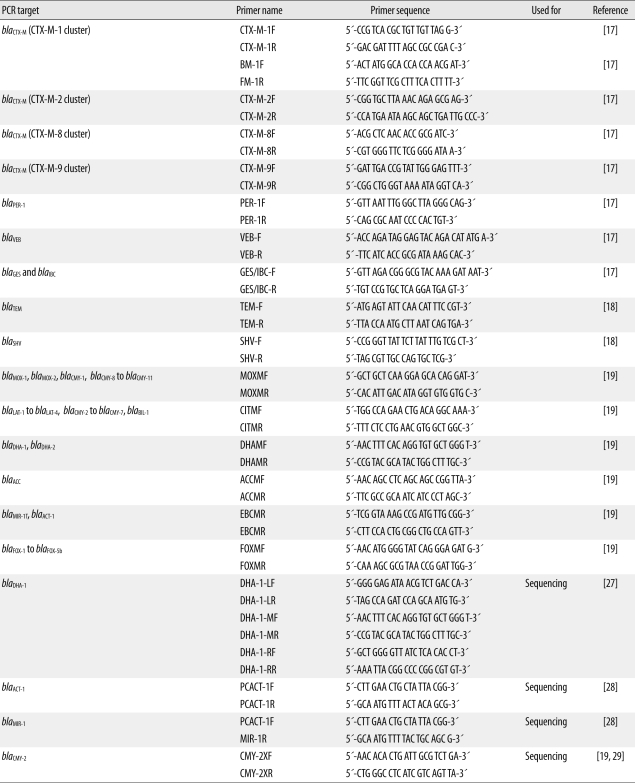
Table 3
Qnr subtype and association with class A beta-lactamase and plasmid-mediated AmpC beta-lactamase




 PDF
PDF ePub
ePub Citation
Citation Print
Print


 XML Download
XML Download