Abstract
Gene manipulation by disrupting important genes using homologous recombination in mammals has provided important insights into their function and development with regard to disease. However, many questions related to the genetic pathways that regulate cellular differentiation and function remain to be clarified. In particular, analysis of genetic pathways that control embryonic skeletal development is often hindered by the disruption of critical genes that function in early embryogenesis, thereby leading to embryonic or perinatal death and thus preventing study of the role of these genes in skeletal development and physiology postnatally. To overcome this problem gene-targeting methods, using site- and time-specific recombination based methods with the Cre/loxP system, have been used to delete particular genes in specific tissues and stages during development. Thus, the generation and characterization of transgenic mice expressing Cre recombinase, under the control of a tissue-specific and stage-specific promoter, has become prerequisite for study of the physiology and homeostasis of specific tissues during a specific time frame. In this report, we introduce the principles and methods of site-specific and time-specific recombination using the Cre/loxP and inducible Cre system, and discuss the potential applications for applying this system to the study of the development and physiology of the skeletal system.
Figures and Tables
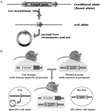 | Fig. 1A, Conditional gene inactiviation by Cre/loxP system. Conditional mice harbor the floxed allele (conditional allele), in which target gene has flanked by two loxP sites. This target gene can be excised and inactivated by Cre recombinase recognizing two loxP sites. B, Transgenic mice expressing Cre recombinase in specific cell types under the control of cell-specific promoter have provided essential tools for studying the function of particular genes in specific tissues. Specific target gene is inactivated by Cre recombinase in specific tissues of the floxed mouse which has normal expression of target gene. |
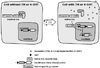 | Fig. 2Inducible Cre system. In the absence of appropriate ligand, the Cre recombinase fused estrogen receptor with mutated ligand binding domain is bound to the heat shock protein (Hsp90) and inhibited from the entering the nucleus. Upon administration and binding of estrogen antagonist, tamoxifen (TM) or 4-hydroxytamoxifen (4-OHT) to the mutant estrogen receptor, Hsp90 dissociates from the CreER complex and allows translocation of activated CreER into the nucleus. In nucleus, CreER recognizes the loxP sites in conditional allele of target genes and mediates recombination. |
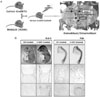 | Fig. 3A, Col1a1-CreERT2; Indicible Cre mouse using 2.3-kb collagen promoter. To determine the ability of Cre recombinase to induce recombination specifically in bone of inducible Cre mouse which has osteoblasts- and odontoblasts-specific collagen promoter, inducible Cre mouse was crossed with the ROSA26 mouse with lacZ reporter. Briefly, the LacZ gene, which is inserted in the ubiquitously expressed ROSA locus, is preceded by a transcriptional stop cassette flanked by loxP sites. Thus, in Col1a1-CreERT2;R26R double transgenic mice, LacZ gene should be expressed in bones when Cre recombinase is activated by 4-OHT administration. Cre recombination activated in a ligand-dependent manner occurred in osteoblasts and odontoblasts under the control of collagen promoter. B, Histological analysis of Col1a1-CreERT2;R26R double transgenic mice. Pups containing the Col1a1-CreERT2;R26R double transgene were injected intraperitoneally with TM or 4-OHT and stained with X-gal. They showed the LacZ expression in all bones and the strong X-gal staining in osteoblasts and odontoblast by histology. No X-gal staining was detected in oil-injected control pups. |
 | Fig. 4Breeding strategy for the conditional gene inactivation using inducible Cre mice. Inducible Cre mice are very useful to study the function of genes in specific tissues or at specific stage of development and after birth. The flox/-;CreER mice harboring the tissue-specific CreER transgene and heterozygous for the floxed and null alleles can be generated. Using flox/-;CreER mice, target genes are inactivated in tissues or cells which Cre recombinase expresses after administration of TM or 4-OHT. |
References
1. Rajewsky K, Gu H, Kuhn R, Betz UA, Muller W, Roes J, Schwenk F. Conditional gene targeting. J Clin Invest. 1996. 98:600–603.
2. Le Y, Sauer B. Conditional gene knockout using Cre recombinase. Methods Mol Biol. 2000. 136:477–485.
3. Sternberg N, Hamilton D. Bacteriophage P1 site-specific recombination. I. Recombination between loxP sites. J Mol Biol. 1981. 150:467–486.
4. Indra AK, Warot X, Brocard J, Bornert JM, Xiao JH, Chambon P, Metzger D. Temporally-controlled site-specific mutagenesis in the basal layer of the epidermis: comparison of the recombinase activity of the tamoxifen-inducible Cre-ER(T) and Cre-ER(T2) recombinases. Nucleic Acids Res. 1999. 27:4324–4327.
5. Feil R, Wagner J, Metzger D, Chambon P. Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochem Biophys Res Commun. 1997. 237:752–757.
6. Leone DP, Genoud S, Atanasoski S, Grausenburger R, Berger P, Metzger D, Macklin WB, Chambon P, Suter U. Tamoxifen-inducible glia-specific Cre mice for somatic mutagenesis in oligodendrocytes and Schwann cells. Mol Cell Neurosci. 2003. 22:430–440.
7. Ducy P, Karsenty G. Two distinct osteoblast-specific cis-acting elements control expression of a mouse osteocalcin gene. Mol Cell Biol. 1995. 15:1858–1869.
8. Rossert J, Eberspaecher H, de Crombrugghe B. Separate cis-acting DNA elements of the mouse pro-alpha-1(I) collagen promoter direct expression of reporter genes to different type I collagen-producing cells in transgenic mice. J Cell Biol. 1995. 129:1421–1432.
9. Dacquin R, Starbuck M, Schinke T, Karsenty G. Mouse alpha-1(I)-collagen promoter is the best known promoter to drive efficient Cre recombinase expression in osteoblast. Dev Dyn. 2002. 224:245–251.
10. Braut A, Kalajzic I, Kalajzic Z, Rowe DW, Kollar EJ, Mina M. Col1a1-GFP transgene expression in developing incisors. Connect Tissue Res. 2002. 43:216–219.
11. Kim JE, Kazuhisa N, de Crombrugghe B. Transgenic mice expressing a ligand-inducible cre recombinase in osteoblasts and odontoblasts: a new tool to examine physiology and disease of postnatal bone and tooth. Am J Pathol. 2004. 165:1875–1882.
12. Soriano P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat Genet. 1999. 21:70–71.
13. Nakashima K, Zhou X, Kunkel G, Zhang Z, Deng JM, Behringer RR, de Crombrugghe B. The novel zinc finger-containing transcription factor osterix is required for osteoblast differentiation and bone formation. Cell. 2002. 108:17–29.
14. Komori T, Yagi H, Nomura S, Yamaguchi A, Sasaki K, Deguchi K, Shimizu Y, Bronson RT, Go YH, Inada M, Sato M, Okamoto R, Kitamura Y, Yoshiki S, Kishimoto T. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell. 1997. 89:755–764.
15. Otto F, Thornell AP, Crompton T, Denzel A, Gilmour KC, Rosewell IR, Stamp GW, Beddington RS, Mundlos S, Olsen BR, Selby PB, Owen MJ. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell. 1997. 89:765–771.
16. Choi JY, Pratap J, Javed A, Zaidi SK, Xing L, Balint E, Dalamangas S, Boyce B, van Wijnen AJ, Lian JB, Stein JL, Jones SN, Stein GS. Subnuclear targeting of Runx/Cbfa/AML factors is essential for tissue-specific differentiation during embryonic development. Proc Natl Acad Sci U S A. 2001. 98:8650–8655.




 PDF
PDF ePub
ePub Citation
Citation Print
Print


 XML Download
XML Download