Abstract
Gaegurin 4 (GGN4), an antimicrobial peptide isolated from a Korean frog, is five times more potent against Gram-positive than Gram-negative bacteria, but has little hemolytic activity. To understand the mechanism of such cell selectivity, we examined GGN4-induced K+ efflux from target cells, and membrane conductances in planar lipid bilayers. The K+ efflux from Gram-positive M. luteus (2.5 μg/ml) was faster and larger than that from Gram-negative E. coli (75 μg/ml), while that from RBC was negligible even at higher concentration (100 μg/ml). GGN4 induced larger conductances in the planar bilayers which were formed with lipids extracted from Gram-positive B. subtilis than in those from E. coli (p< 0.01), however, the effects of GGN4 were not selective in the bilayers formed with lipids from E. coli and red blood cells. Addition of an acidic phospholipid, phosphatidylserine to planar bilayers increased the GGN4-induced membrane conductance (p<0.05), but addition of phosphatidylcholine or cholesterol reduced it (p < 0.05). Transmission electron microscopy revealed that GGN4 induced pore-like damages in M. luteus and dis-layering damages on the outer wall of E. coli. Taken together, the present results indicate that the selectivity of GGN4 toward Gram-positive over Gram-negative bacteria is due to negative surface charges, and interaction of GGN4 with outer walls. The selectivity toward bacteria over RBC is due to the presence of phosphatidylcholine and cholesterol, and the trans-bilayer lipid asymmetry in RBC. The results suggest that design of selective antimicrobial peptides should be based on the composition and topology of membrane lipids in the target cells.
REFERENCES
Agre P., Bennett V. Qualitative and functional analyses of spectrin, ankyrin, band 3, and calmodulin in human red cell membranes. Methods Hematol. 19:95–98. 1988.
Ames GF. Lipids of Salmonella tryphimurium and Escherichia coli: structure and metabolism. J Bacteriol. 95:833–843. 1968.
Balasubramanian K., Schroit AJ. Aminophospholipid asymmetry: a matter of life and death. Annu Rev Physiol. 65:701–734. 2003.

Bishop DG., Rutberg L., Samuelsson B. The chemical composition of the cytoplasmic membrane of Bacillus subtilis. Eur J Biochem. 2:448–453. 1967.

Bishop EA., Bermingham MAC. Lipid composition of Gram-negative bacteria, sensitive and resistant to streptomycin. Antimicrobial Agents Chem. 4:378–379. 1973.

Bloch K. Choleterol: evolution of structure and function. Vance DE, Vance J, editors. ed,. Biochemistry of lipids and membranes. Amsterdam: Elesevier Science Publishers;p. p. 363–382. 1991.
Chi SW., Kim JS., Kim DH., Lee SH., Park YH., Han KH. Solution structure and membrane interaction mode of an antimicrobial peptide gaegurin 4. Biochem Biophys Res Commun. 352:592–597. 2007.

Clark DP., Durell S., Maloy WL., Zasloff M. Ranalexin: a novel antimicrobial peptide from bullfrog (Rana catesbeiana) skin, structurally related to bacterial antibiotic, polymyxin. J Biol Chem. 269:10849–10855. 1994.
Clejan S., Krulwicj TA., Mondrus KR., Sept-Young D. Membrane lipid composition of obligately and facultatively alkalophilic strains of Bacillus spp. J Bacteriol. 168:334–340. 1986.

Conlon JM. Reflections on a systematic nomenclature for antimicrobial peptides from the skins of frogs of the family Ranidae. Peptides. 29:1815–1819. 2008.

Cronan JE., Roy-Vagelos P. Metabolism and function of the membrane phospholipids of Escherichia coli. Biochim Biophys Acta. 165:379–387. 1972.

Daleke DL. Regulation of phospholipid asymmetry in the erythrocyte membrane. Curr Opin Hematol. 15:191–195. 2008.

Dathe M., Schumann M., Wieprecht T., Winkler A., Beyermann M., Krause E., Matsuzaki K., Murase O., Bienert M. Peptide helicity and membrane surface charge modulate the balance of electrostatic and hydrophobic interactions with lipid bilayers and biological membranes. Biochemistry. 35:12612–12622. 1996.

Dowhan W. Molecular basis for membrane phospholipid diversity: why are there so many lipids? Ann Rev Biochem. 66:199–232. 1997.

Eun SY., Jang HK., Han SK., Ryu PD., Lee BJ., Han KH., Kim SJ. A helix-induced oligomeric transition of gaegurin 4, an antimicrobial peptide isolated from a Korean frog. Mol Cells. 21:229–236. 2006.
Gennis RB. The structure and composition of biomembranes. Gennis RB, editor. ed,. Biomembranes, molecular structure and functions. New York: Springer Verlag;p. p. 1–35. 1991.
Gidalevitz D., Ishitsuka Y., Muresan AS., Konovalov O., Waring AJ., Lehrer RI., Lee KY. Interaction of antimicrobial peptide protegrin with biomembranes. Proc Natl Acad Sci USA. 100:6302–6307. 2003.

Kagan B., Selsted ME., Ganz T., Lehrer RI. Antimicrobial defensin peptides form voltage-dependent ion-permeable channels in planar lipid bilayer membranes. Proc Natl Acad Sci USA. 87:210–214. 1990.

Kim HJ., Han SK., Park JB., Baek HJ., Lee BJ., Ryu PD. Gaegurin 4, a peptide antibiotic of frog skin, forms voltage-dependent channels in planar lipid bilayers. J Pept Res. 53:1–7. 1999.

Kim HJ., Kim SS., Lee MH., Lee BJ., Ryu PD. Role of C-terminal heptapeptide in pore-forming activity of antimicrobial agent, gaegurin 4. J Pept Res. 64:151–158. 2004.

Kim KS., Fulton RW. Ultrastructure of Datura stramonium infected with an euphorbia virus suggestive of witefly-transmitted germinivirus. Phytopathology. 74:236–241. 1984.
Lau YH., Caswell AH., Brunschwig J., Baerwald RJ., Garcia M. Lipid analysis and freeze-fracture studies on isolated transverse tubules and sarcoplasmic reticulum subfractions of skeletal muscle. J Biol Chem. 254:540–546. 1979.

Matsumoto K., Kusaka J., Nishibori A., Hara H. Lipid domains in bacterial membranes. Mol Microbiol. 61:1110–1117. 2006.

Matsuzaki K. Why and how are peptide-lipid interactions utilized for self-defense? Magainins and tachyplesins as archetypes. Biochim Biophys Acta. 1462:1–10. 1999.

Matsuzaki K. Control of cell selectivity of antimicrobial peptides. Biochim Biophys Acta. (In press).

Matsuzaki K., Harada M., Handa T., Funakoshi S., Fujii N., Yajima H., Miyajima K. Magainin 1-induced leakage of entrapped calcein out of negatively-charged lipid vesicles. Biochim Biophys Acta. 981:130–134. 1989.

Matsuzaki K., Sugishita K., Fujii N., Miyajima K. Molecular basis for membrane selectivity of an antimicrobial peptide, magainin 2. Biochemistry. 34:3423–3429. 1995.

Morein S., Andersson AS., Rilfors L., Lindblom G. Wild-type Escherichia coli cells regulate the membrane lipid composition in a “Window” between gel and non-lamellar structures. J Biol Chem. 271:6801–6809. 1996.

Morikawa N., Hagiwara K., Nakajima T. Brevinin-1 and brevinin-2, unique antimicrobial peptides from the skin of the frog Rana brevipoda porsa. Biochem Biophys Res Comm. 189:184–190. 1992.
Orlov DS., Nguyen T., Lehrer RI. Potassium release, a useful tool for studying antimicrobial peptides. Microbiol Methods. 49:325–328. 2002.

Park JB., Kim HJ., Ryu PD., Moczydlowski E. Effect of phosphatidylserine on unitary conductance and Ba2+ block of the BK Ca2+-activated K+ channel: re-examination of the surface charge hypothesis. J Gen Physiol. 121:375–398. 2003.
Park S., Son WS., Kim YJ., Kwon AR., Lee BJ. NMR spectroscopic assessment of the structure and dynamic properties of an amphibian antimicrobial peptide (Gaegurin 4) bound to SDS micelles. J Biochem Mol Biol. 40:261–269. 2007.

Park SH., Kim YK., Park JW., Lee BJ., Lee BJ. Solution structure of the antimicrobial peptide gaegurin 4 1H and 15N nuclear magnetic resonance spectroscopy. Eur J Biochem. 267:2695–2704. 2000.
Park JM., Jung JE., Lee BJ. Antimicrobial peptides from the skin of a Korean frog, Rana rugosa. Biochem Biophys Res Comm. 205:948–954. 1994.

Shai Y. From innate immunity to de-novo designed antimicrobial peptides. Curr Pharm Des. 8:715–725. 2002.

Simmaco M., Mignogna G., Barra D., Bossa F. Novel antimicrobial peptides from skin secretion of the European frog, Rana esculenta. FEBA Lett. 324:159–161. 1993.
Fig. 1.
GGN4-induced membrane currents. (A) Induction of membrane conductance by GGN4 (0.3 μg/ml). Current record shows the changes in membrane conductances induced by GGN4 in a lipid bilayer, composed of PE: PS (1: 1), under 200/0 (cis/trans) mM KCl gradient at 0 mV in the recording solution containing 10 mM HEPES-NMDG (pH 7.2). (B and C) Typical records showing unitary conductances of GGN4 (0.03 μg/ml)-induced pores at symmetrical 100 mM KCl in acidic (PE: PS=3: 7, B) and neutral lipid membranes (PE 100%, C) at 30 mV.
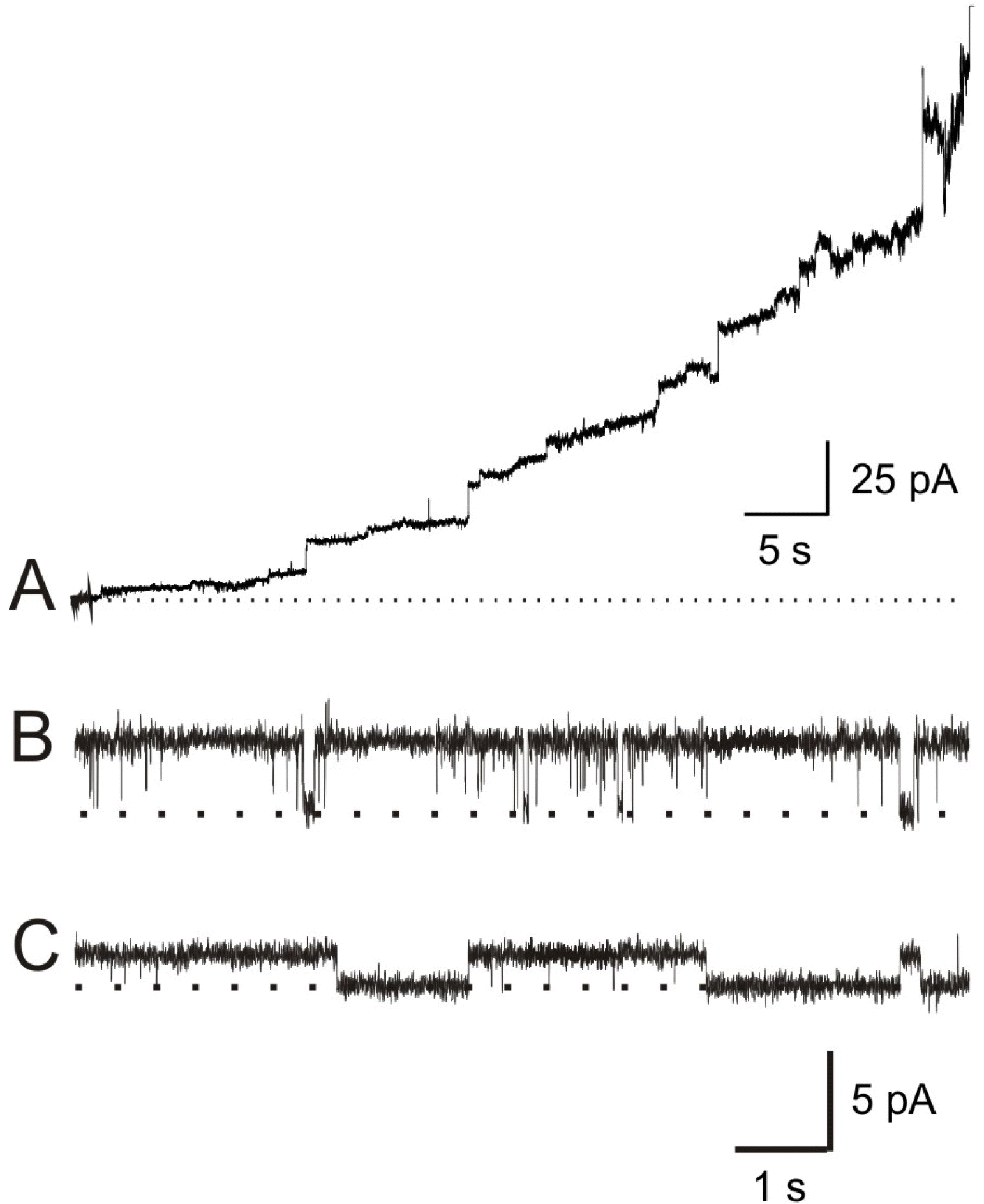
Fig. 2.
Effects of GGN4 on K+ efflux from bacteria and RBC in the solution containing 10 mM HEPES-NaOH (pH 7.0) and 0.15 M NaCl. (A-C) K+ efflux from M. luteus (A), E. coli (B), and RBC (1%, v/v) in the presence of GGN4 (C). (D) Time courses of K+ efflux (Δ) and cell viability (○) after application of GGN4 (12.5 μg/ml) to M. luteus suspension. The extent of K+ efflux was normalized to the total K+ efflux obtained after treatment with 0.5% Triton X-100.
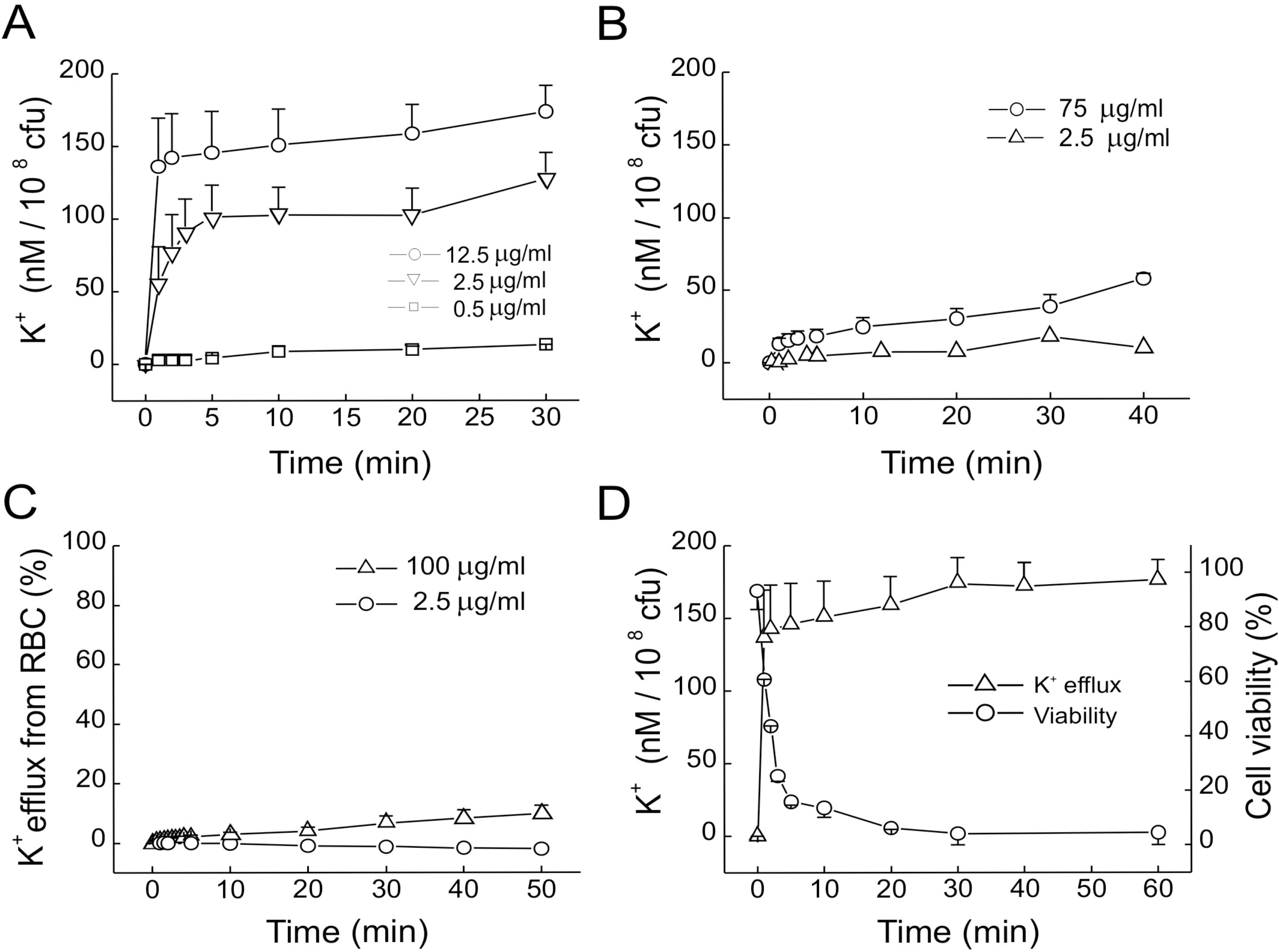
Fig. 3.
Pore-forming activity of GGN 4 in the membranes formed with lipids extracted from G(+), Gram-negative bacteria and RBC in symmetric 25 mM KCl. (A) Current-voltage relation of GGN4-induced channels in the membranes formed with the extracted lipids in response to a ramp pulse (0.03 μg/ml in E. coli lipid membrane and 0.1 μg/ml in B. subtilis lipid membrane). (B) Mean slope conductances of the membranes formed with lipids extracted from B. subtilis, E. coli and RBC. Bars represent standard error of means. n=6.
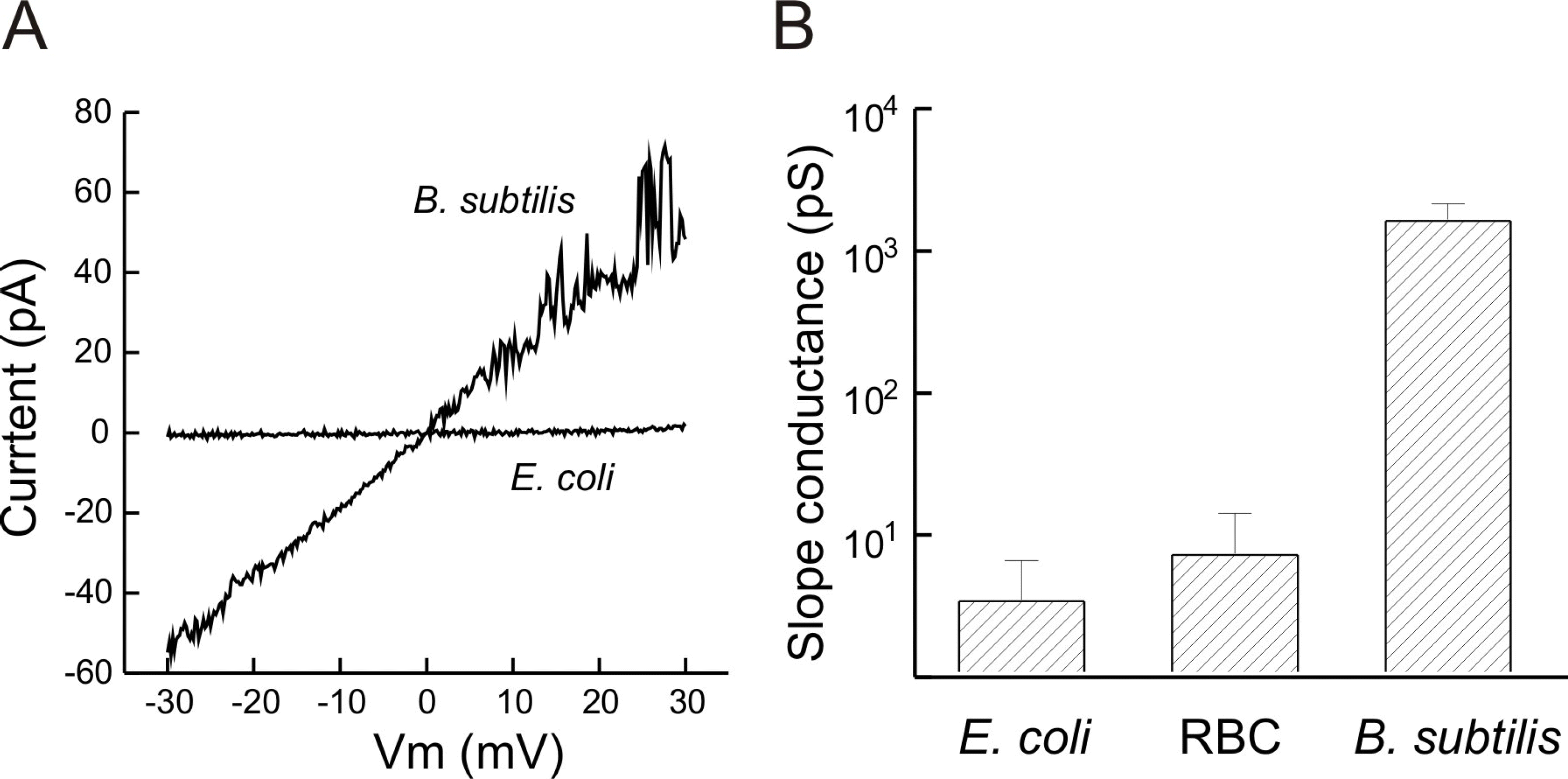
Fig. 4.
Effects of addition of PC, PS and cholesterol to the lipid bilayers on GGN4-induced conductances. Slope conductances of the membrane at each GGN4 concentration were estimated from the current-voltage relations as shown in Fig. 3. Solid lines are drawn by the linear regression of double logarithmic concentration-conductance relations. Respective slopes of log space (conductance, pS) vs. log. (concentration, μg/ml) are 0.391, 0.546, 0.738 and 0.704 for PE, PE: PC, PE: PC: PS, and PE: PC: PS: CS membranes, respectively. PE, 100% phosphatidylethanolamine; PE: PC, 80% phosphatidylethanolamine and 20% phosphatidylcholine; PE: PC: PS, 80% phosphatidylethanolamine, 10% phosphatidylcholine and 10% phosphatidylserine; PE: PC: PS: CS, 50% phosphatidylethanolamine, 10% phosphatidylcholine, 10% phosphatidylserine, and 30% cholesterol. Symbols and bars represent the means and error bars of slope conductances, respectively, measured from 3~6 bilayers, except the data point at 0.3 μg/ml in PE: PC: PS: CS membrane, where n=2.
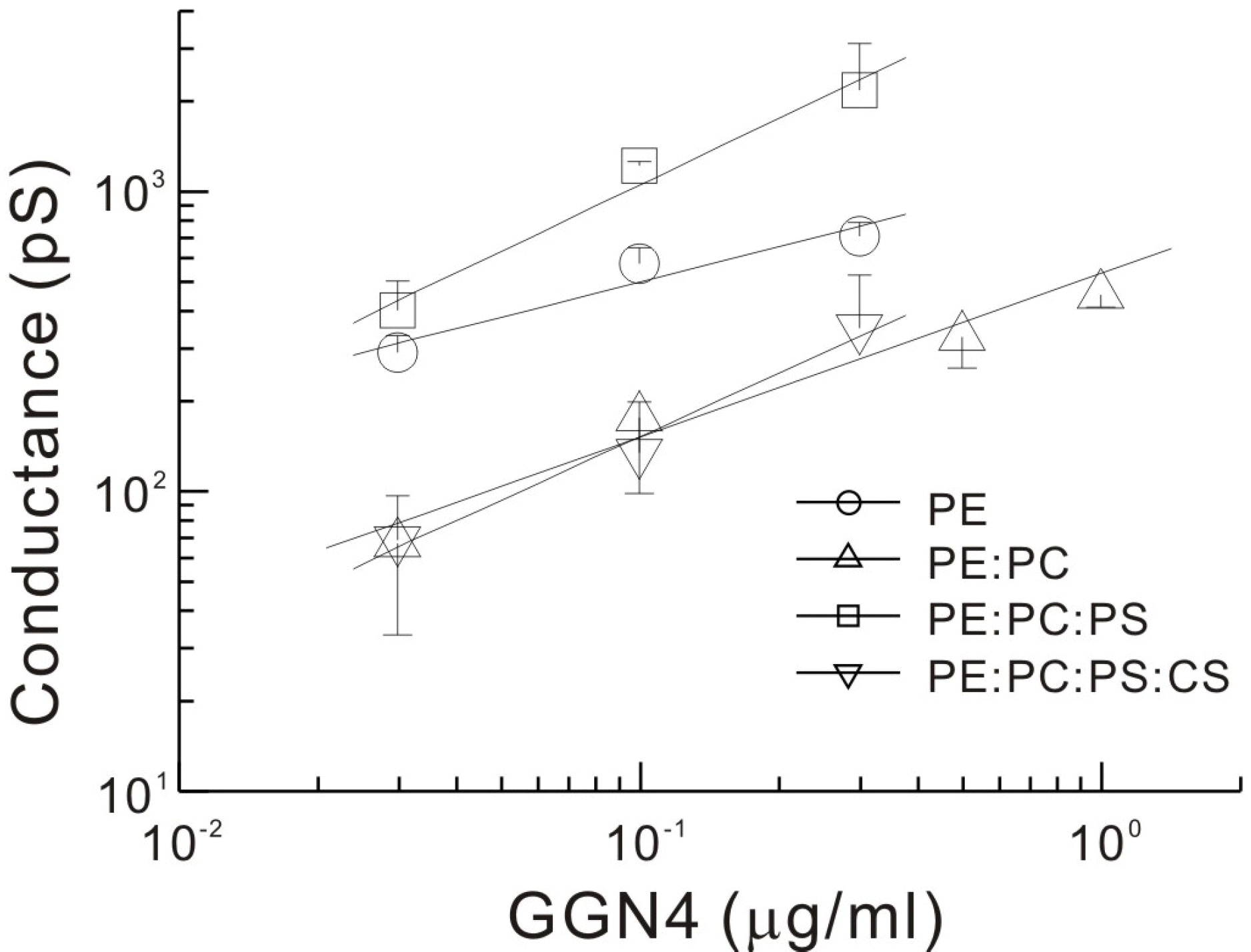
Fig. 5.
Transmission electron micrographs of M. luteus and E. coli treated with GGN4 at 37°C for 30 min. M. luteus and E. coli cells (108 cfu/ml) were incubated with 2.5 and 75 μg/ml GGN4, respectively. (A~C) Transmission electron micrographs of M. luteus untreated (A) and treated with GGN4 (B and C). Note the pores (marked by arrows) on the bacterial membranes (B). One of the pores shown in (B) was illustrated at higher magnification to show cytoplasm leaking out through the pore of a diameter of about 55 nm (marked by ∗, C). (D~F) Transmission electron micrographs of E. coli untreated (D) and treated with GGN4 (E and F). Note the layers of outer wall debris were peeled off from the damaged membranes (arrows) and a space developed between outer wall and inner membrane of E. coli at a damaged site (E, arrow). The bars represent 0.5 μm for (A), (B), (D) and (E), and 0.1 μm for (C) and (F).





 PDF
PDF ePub
ePub Citation
Citation Print
Print


 XML Download
XML Download