SUMMARY
The genetic basis of autoimmune thyroid disease (AITD), which includes Graves' disease (GD) and Hashimoto's thyroiditis (HT), is thought to be polygenic. The genetic factors have been shown by twin studies and familial aggregation, including clustering of GD and HT within the same families. While the genetic basis of AITD is unclear, it is believed to be polygenic. Identification of genetic factors will contribute to not only the understanding of pathogenesis of AITD, but also the prevention and treatment of the diseases. Several genetic factors associated with AITD susceptibility have been tentatively identified. Some are molecules related to immune responses including the human leukocyte antigen (HLA) and the cytotoxic T lymphocyte associated-4 (CTLA-4) genes. Others are those related to thyroid-specific autoantigens including the thyrotropin receptor (TSHR) gene. As part of a genome scan to locate familial GD and HT genes, other additional chromosomal locations including chromosome 8q and 5q have been reported in a Japanese population. As the next step, ZFAT (zinc-finger gene in AITD susceptibility region) gene was identified as one of the susceptibility genes in 8q23-q24. In addition, we found that a particular allele of intron 7 or near this region of the TSHR gene may contribute to GD susceptibility. Here, I introduce our recent studies on genetics of AITD.
INTRODUCTION
Autoimmune thyroid disease (AITD) is a multifactorial disease with a significant genetic and environmental component (Fig. 1). These factors cause the impairment of self tolerance to thyroid autoantigens and eventually develop AITD. The genetic factors have been shown by twin studies and familial aggregation, including clustering of Graves' disease (GD) and Hashimoto's thyroiditis (HT) within the same families (Fig. 2)[1~3]. In GD, relative risk of the first-degree relatives has been reported between 16 to 26 in Caucasians. We found that, it was 19-42 in Japanese population, the relative risk of siblings, λs, was approximately half of them, 7.5-15. More obviously, GD develops much more often in monozygotic twins than dizygotic twins. Also in animal models, genetic components were suggested in both experimental and spontaneous models. Their genes include major histocompatibility complex (MHC) and other genes. Thus, existence of genetic factors in AITD is strongly suggested.
We expect that genetic study of AITD will provide us several valuable information. First of all, genetic study definitely contributes to the clarification of pathogenesis of AITD. It will also be helpful to investigate environmental factors. Secondly, genetic study will be useful for estimation and prevention of disease development. Genetic difference may determine susceptibility of drug- or food-induced thyroid dysfunction and postpartum hypo- or hyperthyroidism. Thirdly, genetic study will provide valuable information on treatment. It is well known that response to anti-thyroid drug is variable among Graves' patients, and that side effects of anti-thyroid drugs are not rare and sometimes very severe. Genetic information will be helpful to obtain good response and to avoid side effects of drug treatment.
There are two approaches to identify genetic factors, candidate gene approach and genome-wide screening. From a viewpoint of DNA samples, two methods, case-control study and family study are commonly used. By using these methods and subjects, susceptibility genes of AITD have been searched widely over the world. Indeed, several genetic factors associated with AITD have been tentatively identified by candidate-gene and genome-scanning approaches. In the candidate approach, genes related to immune responses and thyroid-specific autoantigens were studied. Then, the human leukocyte antigen (HLA), cytotoxic T lymphocyte associated-4 (CTLA-4), interleukin-1 receptor antagonist, interleukin-4 (IL-4), thyrotropin receptor (TSHR) and thyroglobulin (Tg) genes were raised[4~15]. In the genome-scanning approach, other chromosomal regions, such as 14q31 (GD-1), 20q11.2 (GD-2), Xq21.33-22 (GD-3), 13q32 (HT-1), 12q22, 18q21 (IDDM6), Xp11, 8q23-q24, 5q31-q33[4,5,7,8,10~16]. Evidence for AITD genes, however, varies among researchers or populations, and no consensus has been obtained.
Fig. 3 illustrates possible susceptibility genes and loci for AITD found in Japanese population. Underline shows genes found by candidate gene approach and others shows loci found by genome-wide screening. All genes or loci have also been raised as susceptibility genes for AITD in Caucasians, although no consensus has been achieved. Concerning CTLA-4 and HLA, a lot of evidence has already accumulated. First of all, I briefly refer to these two genes.
CTLA-4 and HLA genes
CTLA-4 is co-expressed with CD28 on activated T-cells and interacts with B7 on antigen-presenting cells to stimulate T-cell proliferation. Several polymorphisms of CTLA-4 are known (Fig. 4). And their associations with AITD have been demonstrated in different populations by many researchers including Koreans and ourselves[4,17]. Further, for two SNPs, 49A/G SNP and a SNP located 3' downstream of exon 4, functional analyses were attempted [18,19].
In 49A/G SNP, which is located in the signal sequence, G/G genotype is associated with AITD[7]. Reflecting this, proliferation of peripheral T lymphocytes was higher in patients with G/G genotype than in other genotypes[18]. Further, CTLA-4 blockade of T cell proliferation was lower in G/G genotype than in other genotypes, suggesting that CTLA-4 function was lower and autoimmune response was more activated in patients with G/G genotype. In another SNP of CTLA-4, CT60 SNP, G/G genotype of this SNP was also associated with AITD[19]. There are two forms of CTLA-4, full-length form and soluble form. Soluble form is thought be more important for CTLA-4 function. The expression of soluble form was less in patients with G/G genotype than in other genotypes. This result also suggested that CTLA-4 function was lower and autoimmune response was more activated in patients with G/G genotype. Thus, CTLA-4 is one of molecules studied in the most advanced level as a susceptibility gene of AITD.
HLA is a key molecule to discriminate self from non-self and essential to immunotolerance mechanism. Therefore, it is no doubt that HLA is one of genetic factors for autoimmune diseases including AITD. In fact, strong association of DR3 with AITD was demonstrated in Caucasians[8,20]. Also in Asians, association of HLA with AITD was reported. For example, in Japanese, association of DPB10501, A2 and DQ0501 was found[20]. However, there remain several problems and questions for HLA in Asian population. Because association of HLA in Asians is much weaker than that in Caucasians, it is doubtful whether HLA is a major gene of AITD in Asians or not. No definite consensus for allele types of HLA which are associated with AITD has been achieved yet in Asians. Finally, functional analysis has not thoroughly been performed. There several questions and problems that remain to be answered.
Genome-wide Screening: 5q31-q33 & 8q23-q24
Sakai et al. carried out affected sib-pair linkage analysis using 113 Japanese AITD families[16]. Most families had two affected siblings. Some families were mixed with GD and HT patients. They searched susceptibility loci genome-widely using 392 microsatellite markers, which are about 10 cM-spaced. They found suggestive linkage to AITD at two loci, which gave high LOD scores more than 3. One was 5q31-q33, in which LOD scores to AITD and GD were 3.1 and 1.5, respectively. The other locus was 8q23-q24, in which LOD scores to HT and AITD were 3.8 and 2.3, respectively.
5q31-q33 region is intriguing because cytokine cluster including IL3, GM-CSF, IL13, IL4, and IL9 are located (Fig. 5). Other genes related to immunoregulation including interleukin related factor 1 (IRF-1), CD14 and beta-2 adrenergic receptor are also present in this locus. In fact, this locus was raised as a possible susceptibility locus for several autoimmune and allergic diseases including atopy and asthma[21~24]. Also in AITD, association of several SNPs of genes located in this locus including IL13, IL4 and IRF-1 was reported[9,24~27]. Therefore, we attempted to narrow this locus to identify a susceptibility gene of AITD by using many genetic markers located in this locus[28]. However, so far, we have not succeeded. This is possibly because the susceptibility gene in this locus is too minor to identify with limited genetic markers and subjects. Alternatively, because many immune-related genes are present in this locus, not a single but multiple genes in this locus may contribute and interact each other. In this case, it is difficult to identify a gene by searching a strong association or linkage.
In contrast, Shirasawa et al. succeeded in finding a susceptibility gene in 8q23-q24 locus[29]. This region is interesting because thyroglobulin gene is located. They attempted to narrow 21Mb region by using 169 microsatellite markers, which were 120kb-spaced. They found a deep trough of p values in association study. This position was slightly different from that of Tg gene. Then, they further tried to narrow 400kb region using 36 SNPs, which were 11kb-spaced. Again, they found a deep trough of p-value. Finally, they identified a gene located in this region, named zinc-finger gene in AITD susceptibility region, ZFAT. ZFAT genome consists of 19 exons and has five splicing variants. A truncated form, TR-ZFAT has a stop codon in exon 9. A small anti-sense form of ZFAT, SAS-ZFAT has only two exons and is transcribed in a reverse direction. As the name, 'zinc-finger' suggested, ZFAT is presumed to be a transcription factor. The expression of ZFAT was ubiquitous. TR-ZFAT was also ubiquitously present, although less than ZFAT. In contrast, SAS-ZFAT was expressed predominantly in B lymphocytes. Interestingly, a SNP in Exon9b, SNP10 affected the expression level of SAS-ZFAT. The expression of SAS-FAT was decreased in T/T genotype. In fact, strong association of T allele of this SNP with GD and HT had been demonstrated in case-control study. The T allele of this SNP was significantly increased in AITD than in control. This SNP was located in the putative promoter region of SAS-ZFAT. In the luciferase assay, the promotor activities were different by the genotype of this SNP; the activity was lower in T allele than in A allele. In addition, in gel shift assay, the presence of a binding protein to T allele was indicated. Further, they examined whether SAS-ZFAT transcript itself affects the expression level of TR-ZFAT or not. In a co-transfection experiments with TR-ZFAT and SAS-ZFAT expression vectors in HEK293 cells, expression of TR-ZFAT was reduced by expression of SAS-ZFAT, while empty or unrelated (Mig6-30-UTR) vector did not change. Expression of ZFAT was not changed by SAS-ZFAT. The stability of TR-ZFAT mRNA was not different by the genotypes of this SNP. Thus, TR-ZFAT expression seems to be regulated by the SAS-ZFAT transcript. And this SNP might be related to the susceptibility of AITD (Fig. 6).
Candidate gene approach: TSHR
Because TSHR is a key autoantigen for GD, several groups have already investigated association or linkage of TSHR gene polymorphism with AITD since 1995[4,10,30~32]. They used SNPs including codon 52 and codon 727, and mirosatellite markers for association and linkage studies in Caucasians or Asian population. Some were positive results, but others were negative. The consensus has not been obtained yet. In the previous studies using SNPs, there are several problems. The number of SNPs used was small. Exon SNPs only were used. The frequency of minor allele of SNP was low, less than 10%. Linkage disequilibrium (LD) analysis was poorly performed.
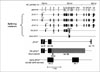
To solve these problems, we systemically searched Japanese SNPs in J-SNP data base, including intron SNPs[33]. We selected SNP with the minor allele frequency of not less than 10%. We performed full LD analysis of SNPs within TSHR gene. Figure 7 shows the localization of 11 SNPs which we used for the following study. They cover from exon 1 to 3' downstream region adjacent to exon 10. They spaced 10~50 kb apart. Because a SNP in intron 7 demonstrated a strong association with GD, additional SNPs were searched in J-SNP. We recruited ~400 Japanese AITD subjects including 250 GD and ~151 HT, and 237 controls. Genotyping was analyzed by SNaP shot and direct DNA sequencing. All SNPs were not deviated from Hardy-Weinberg equilibrium. Several SNPs located in or near intron 7 showed a strong association with GD, but not HT. For example, p value of this SNP (JST022302) was 0.0004. In HT, only weak association was observed in intron 1 SNP. Next, we examined LD of these SNPs using two methods. First of all, based on D' absolute values, two blocks appeared to be present in the TSHR gene. One block extends from intron 5 to exon 8, and the other from intron 8 to 3'UTR. The first block was divided into three blocks based on r-square values. All polymorphisms present in block 3 showed strong association with GD, while SNPs in other blocks showed only limited evidence for association. Therefore, we performed haplotype analysis using block 3 SNPs. In case-control haplotype association analysis using 8 polymorphisms in block 3, only two common haplotypes were observed. The haplotype of all major allele was significantly increased in GD, but not HT, while that of all minor alleles was significantly decreased. Thus, haplotype analysis confirmed the result of the single SNP analysis. These findings suggest that a particular allele of intron 7 or near this region of the TSHR gene may contribute to GD susceptibility.
Very recently, Dechairo et al. reported association of TSHR gene SNP with GD[34]. They found 3 blocks across the TSHR region. A haplotype across first two blocks showed a strong association with GD, but not HT. Since SNPs and subjects that they used were completely different from ours, it is impossible to compare their results directly with ours. In fact, the positions of blocks that they found were different from ours. However, it is certain that TSHR is emerging as a GD-specific susceptibility gene. Further fine mapping and functional analysis are necessary to identify the etiological variants.
EPILOGUE
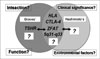
I showed several possible susceptibility genes or loci of AITD, including HLA, CTLA-4, ZFAT, 5q31-33 and TSHR (Fig. 8). All except TSHR are common to GD and HT. TSHR gene is, so far, the only disease-specific gene. However, we have only limited information on susceptibility genes of AITD. Other susceptibility genes may exist. Particularly, HT-specific genes have not yet been found. Interactions between genes have not fully been analyzed. Functional analysis of these genes has not been thoroughly performed, either. We do not know clinical significance of these genes in AITD. Finally, we do not know the interaction with environmental factors, either. We certainly need future vigorous work to answer these questions.




 PDF
PDF ePub
ePub Citation
Citation Print
Print









 XML Download
XML Download